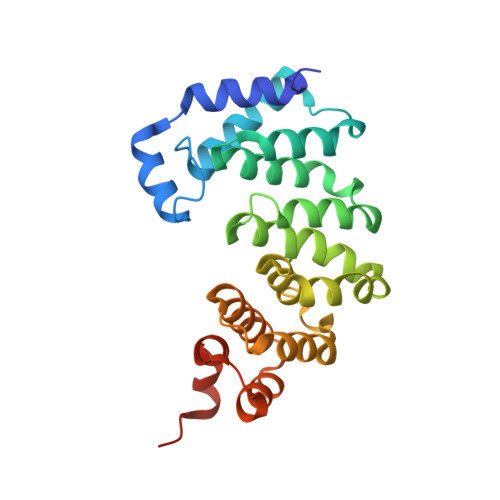
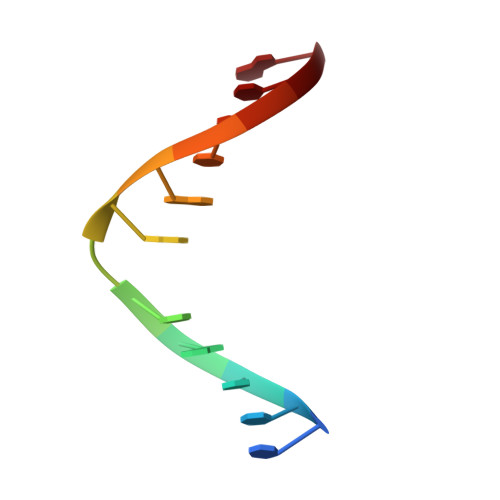
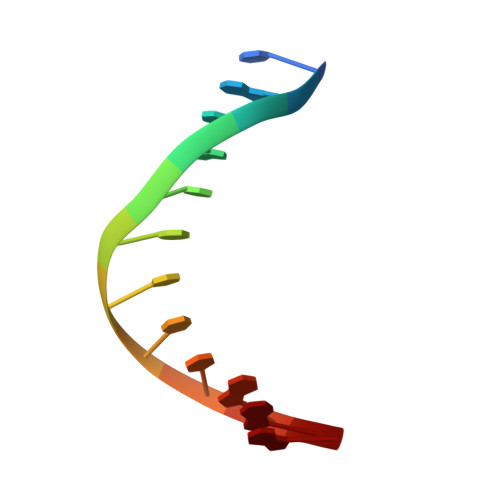
A Catalytic Role for C-H/ pi Interactions in Base Excision Repair by Bacillus cereus DNA Glycosylase AlkD.
Parsons, Z.D., Bland, J.M., Mullins, E.A., Eichman, B.F.(2016) J Am Chem Soc 138: 11485-11488
- PubMed: 27571247 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.6b07399
- Primary Citation Related Structures:
5KUB - PubMed Abstract:
DNA glycosylases protect genomic integrity by locating and excising aberrant nucleobases. Substrate recognition and excision usually take place in an extrahelical conformation, which is often stabilized by π-stacking interactions between the lesion nucleobase and aromatic side chains in the glycosylase active site. Bacillus cereus AlkD is the only DNA glycosylase known to catalyze base excision without extruding the damaged nucleotide from the DNA helix. Instead of contacting the nucleobase itself, the AlkD active site interacts with the lesion deoxyribose through a series of C-H/π interactions. These interactions are ubiquitous in protein structures, but evidence for their catalytic significance in enzymology is lacking. Here, we show that the C-H/π interactions between AlkD and the lesion deoxyribose participate in catalysis of glycosidic bond cleavage. This is the first demonstration of a catalytic role for C-H/π interactions as intermolecular forces important to DNA repair.
- Department of Biological Sciences and Center for Structural Biology, Vanderbilt University , Nashville, Tennessee 37232, United States.
Organizational Affiliation: