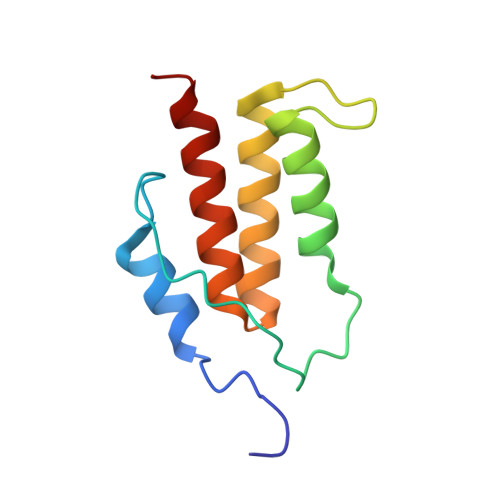
Structure of the C-Terminal Helical Repeat Domain of Eukaryotic Elongation Factor 2 Kinase.
Will, N., Piserchio, A., Snyder, I., Ferguson, S.B., Giles, D.H., Dalby, K.N., Ghose, R.(2016) Biochemistry 55: 5377-5386
- PubMed: 27571275 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b00711
- Primary Citation Related Structures:
5KS5 - PubMed Abstract:
Eukaryotic elongation factor 2 kinase (eEF-2K) phosphorylates its only known physiological substrate, elongation factor 2 (eEF-2), which reduces the affinity of eEF-2 for the ribosome and results in an overall reduction in protein translation rates. The C-terminal region of eEF-2K, which is predicted to contain several SEL-1-like helical repeats (SLRs), is required for the phosphorylation of eEF-2. Using solution nuclear magnetic resonance methodology, we have determined the structure of a 99-residue fragment from the extreme C-terminus of eEF-2K (eEF-2K627-725) that encompasses a region previously suggested to be essential for eEF-2 phosphorylation. eEF-2K627-725 contains four helices, of which the first (αI) is flexible, and does not pack stably against the ordered helical core formed by the last three helices (αII-αIV). The helical core is structurally similar to members of the tetratricopeptide repeat (TPR) family that includes SLRs. The two penultimate helices, αII and αIII, comprise the TPR, and the last helix, αIV, appears to have a capping function. The eEF-2K627-725 structure illustrates that the C-terminal deletion that was shown to abolish eEF-2 phosphorylation does so by destabilizing αIV and, therefore, the helical core. Indeed, mutation of two conserved C-terminal tyrosines (Y712A/Y713A) in eEF-2K previously shown to abolish eEF-2 phosphorylation leads to the unfolding of eEF-2K627-725. Preliminary functional analyses indicate that neither a peptide encoding a region deemed crucial for eEF-2 binding nor isolated eEF-2K627-725 inhibits eEF-2 phosphorylation by full-length eEF-2K. Taken together, our data suggest that the extreme C-terminal region of eEF-2K, in isolation, does not provide a primary docking site for eEF-2.
- Department of Chemistry and Biochemistry, The City College of New York , New York, New York 10031, United States.
Organizational Affiliation: