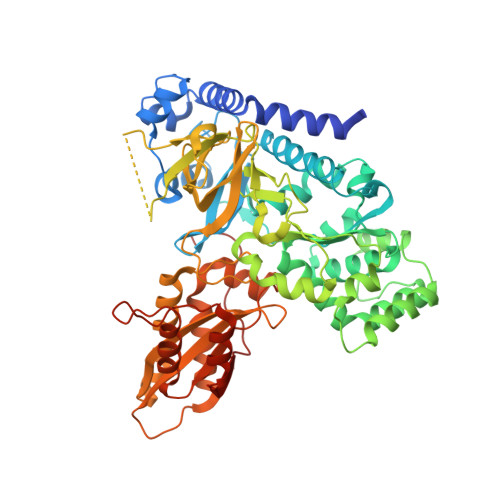
Arabidopsis thaliana GH3.5 acyl acid amido synthetase mediates metabolic crosstalk in auxin and salicylic acid homeostasis.
Westfall, C.S., Sherp, A.M., Zubieta, C., Alvarez, S., Schraft, E., Marcellin, R., Ramirez, L., Jez, J.M.(2016) Proc Natl Acad Sci U S A 113: 13917-13922
- PubMed: 27849615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1612635113
- Primary Citation Related Structures:
5KOD - PubMed Abstract:
In Arabidopsis thaliana, the acyl acid amido synthetase Gretchen Hagen 3.5 (AtGH3.5) conjugates both indole-3-acetic acid (IAA) and salicylic acid (SA) to modulate auxin and pathogen response pathways. To understand the molecular basis for the activity of AtGH3.5, we determined the X-ray crystal structure of the enzyme in complex with IAA and AMP. Biochemical analysis demonstrates that the substrate preference of AtGH3.5 is wider than originally described and includes the natural auxin phenylacetic acid (PAA) and the potential SA precursor benzoic acid (BA). Residues that determine IAA versus BA substrate preference were identified. The dual functionality of AtGH3.5 is unique to this enzyme although multiple IAA-conjugating GH3 proteins share nearly identical acyl acid binding sites. In planta analysis of IAA, PAA, SA, and BA and their respective aspartyl conjugates were determined in wild-type and overexpressing lines of A thaliana This study suggests that AtGH3.5 conjugates auxins (i.e., IAA and PAA) and benzoates (i.e., SA and BA) to mediate crosstalk between different metabolic pathways, broadening the potential roles for GH3 acyl acid amido synthetases in plants.
- Department of Biology, Washington University in St. Louis, St. Louis, MO 63130.
Organizational Affiliation: