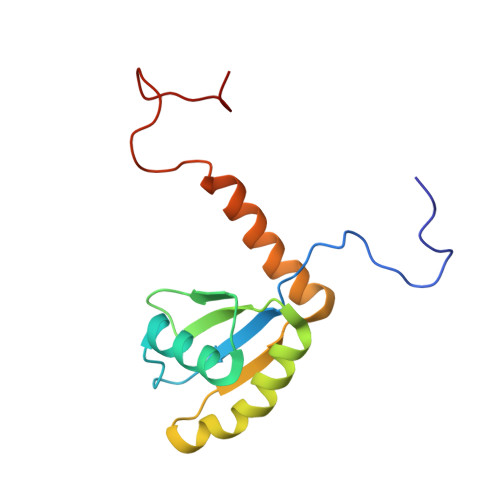
hLARP7 C-terminal domain contains an xRRM that binds the 3' hairpin of 7SK RNA.
Eichhorn, C.D., Chug, R., Feigon, J.(2016) Nucleic Acids Res 44: 9977-9989
- PubMed: 27679474 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkw833
- Primary Citation Related Structures:
5KNW - PubMed Abstract:
The 7SK small nuclear ribonucleoprotein (snRNP) sequesters and inactivates the positive transcription elongation factor b (P-TEFb), an essential eukaryotic mRNA transcription factor. The human La-related protein group 7 (hLARP7) is a constitutive component of the 7SK snRNP and localizes to the 3' terminus of the 7SK long noncoding RNA. hLARP7, and in particular its C-terminal domain (CTD), is essential for 7SK RNA stability and assembly with P-TEFb. The hLARP7 N-terminal La module binds and protects the 3' end from degradation, but the structural and functional role of its CTD is unclear. We report the solution NMR structure of the hLARP7 CTD and show that this domain contains an xRRM, a class of atypical RRM first identified in the Tetrahymena thermophila telomerase LARP7 protein p65. The xRRM binds the 3' end of 7SK RNA at the top of stem-loop 4 (SL4) and interacts with both unpaired and base-paired nucleotides. This study confirms that the xRRM is general to the LARP7 family of proteins and defines the binding site for hLARP7 on the 7SK RNA, providing insight into function.
- Department of Chemistry and Biochemistry, P.O. Box 951569, University of California, Los Angeles, CA 90095-1569, USA.
Organizational Affiliation: