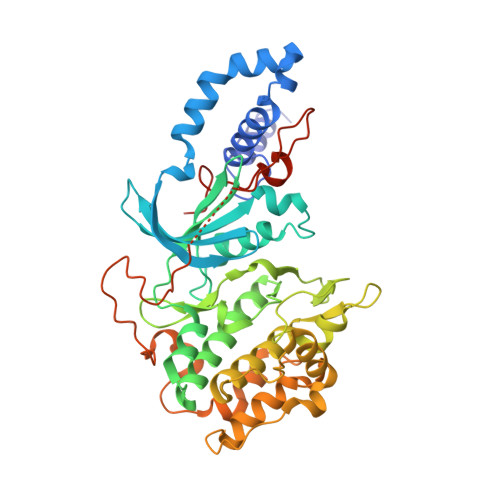
ROCK inhibitors 3: Design, synthesis and structure-activity relationships of 7-azaindole-based Rho kinase (ROCK) inhibitors
Bandarage, U.K., Cao, J., Come, J.H., Court, J.J., Gao, H., Jacobs, M.D., Marhefka, C., Nanthakumar, S., Green, J.(2018) Bioorg Med Chem Lett