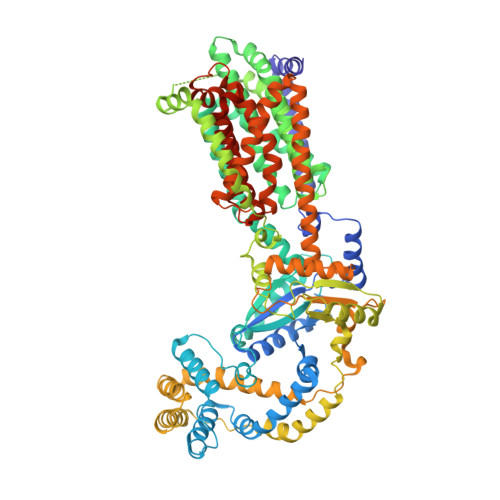
Crystal structures of the Burkholderia multivorans hopanoid transporter HpnN.
Kumar, N., Su, C.C., Chou, T.H., Radhakrishnan, A., Delmar, J.A., Rajashankar, K.R., Yu, E.W.(2017) Proc Natl Acad Sci U S A 114: 6557-6562
- PubMed: 28584102 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1619660114
- Primary Citation Related Structures:
5KHN, 5KHS - PubMed Abstract:
Strains of the Burkholderia cepacia complex (Bcc) are Gram-negative opportunisitic bacteria that are capable of causing serious diseases, mainly in immunocompromised individuals. Bcc pathogens are intrinsically resistant to multiple antibiotics, including β-lactams, aminoglycosides, fluoroquinolones, and polymyxins. They are major pathogens in patients with cystic fibrosis (CF) and can cause severe necrotizing pneumonia, which is often fatal. Hopanoid biosynthesis is one of the major mechanisms involved in multiple antimicrobial resistance of Bcc pathogens. The hpnN gene of B. multivorans encodes an integral membrane protein of the HpnN family of transporters, which is responsible for shuttling hopanoids to the outer membrane. Here, we report crystal structures of B. multivorans HpnN, revealing a dimeric molecule with an overall butterfly shape. Each subunit of the transporter contains 12 transmembrane helices and two periplasmic loops that suggest a plausible pathway for substrate transport. Further analyses indicate that HpnN is capable of shuttling hopanoid virulence factors from the outer leaflet of the inner membrane to the periplasm. Taken together, our data suggest that the HpnN transporter is critical for multidrug resistance and cell wall remodeling in Burkholderia .
- Department of Chemistry, Iowa State University, Ames, IA 50011.
Organizational Affiliation: