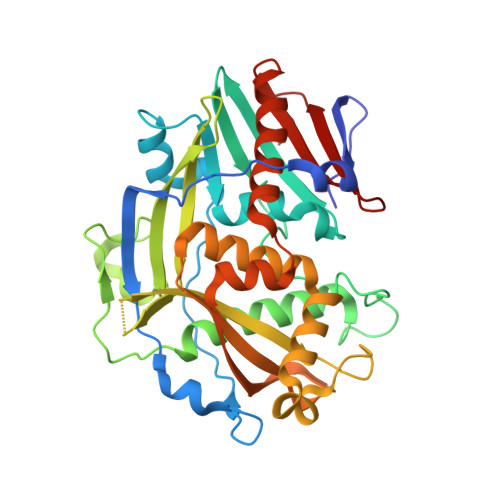
eIF3d is an mRNA cap-binding protein that is required for specialized translation initiation.
Lee, A.S., Kranzusch, P.J., Doudna, J.A., Cate, J.H.(2016) Nature 536: 96-99
- PubMed: 27462815 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature18954
- Primary Citation Related Structures:
5K4B, 5K4C, 5K4D - PubMed Abstract:
Eukaryotic mRNAs contain a 5′ cap structure that is crucial for recruitment of the translation machinery and initiation of protein synthesis. mRNA recognition is thought to require direct interactions between eukaryotic initiation factor 4E (eIF4E) and the mRNA cap. However, translation of numerous capped mRNAs remains robust during cellular stress, early development, and cell cycle progression despite inactivation of eIF4E. Here we describe a cap-dependent pathway of translation initiation in human cells that relies on a previously unknown cap-binding activity of eIF3d, a subunit of the 800-kilodalton eIF3 complex. A 1.4 Å crystal structure of the eIF3d cap-binding domain reveals unexpected homology to endonucleases involved in RNA turnover, and allows modelling of cap recognition by eIF3d. eIF3d makes specific contacts with the cap, as exemplified by cap analogue competition, and these interactions are essential for assembly of translation initiation complexes on eIF3-specialized mRNAs such as the cell proliferation regulator c-Jun (also known as JUN). The c-Jun mRNA further encodes an inhibitory RNA element that blocks eIF4E recruitment, thus enforcing alternative cap recognition by eIF3d. Our results reveal a mechanism of cap-dependent translation that is independent of eIF4E, and illustrate how modular RNA elements work together to direct specialized forms of translation initiation.