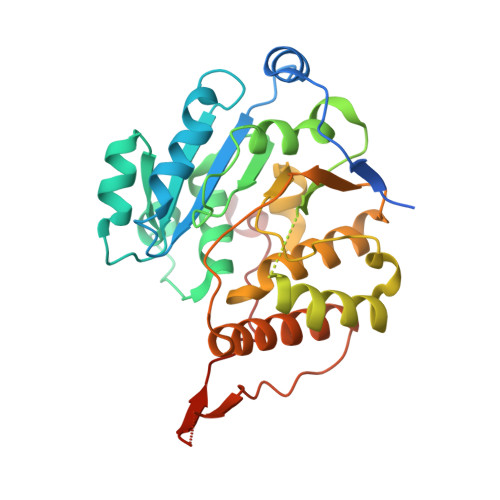
Structural Snapshots and Loop Dynamics along the Catalytic Cycle of Glycosyltransferase GpgS.
Albesa-Jove, D., Romero-Garcia, J., Sancho-Vaello, E., Contreras, F.X., Rodrigo-Unzueta, A., Comino, N., Carreras-Gonzalez, A., Arrasate, P., Urresti, S., Biarnes, X., Planas, A., Guerin, M.E.(2017) Structure 25: 1034-1044.e3
- PubMed: 28625787 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2017.05.009
- Primary Citation Related Structures:
5JQX, 5JSX, 5JT0, 5JUC, 5JUD - PubMed Abstract:
Glycosyltransferases (GTs) play a central role in nature. They catalyze the transfer of a sugar moiety to a broad range of acceptor substrates. GTs are highly selective enzymes, allowing the recognition of subtle structural differences in the sequences and stereochemistry of their sugar and acceptor substrates. We report here a series of structural snapshots of the reaction center of the retaining glucosyl-3-phosphoglycerate synthase (GpgS). During this sequence of events, we visualize how the enzyme guides the substrates into the reaction center where the glycosyl transfer reaction takes place, and unveil the mechanism of product release, involving multiple conformational changes not only in the substrates/products but also in the enzyme. The structural data are further complemented by metadynamics free-energy calculations, revealing how the equilibrium of loop conformations is modulated along these itineraries. The information reported here represent an important contribution for the understanding of GT enzymes at the molecular level.
- Structural Biology Unit, CIC bioGUNE, Bizkaia Technology Park, Ed. 801A, 48160 Derio, Spain; Unidad de Biofísica, Centro Mixto Consejo Superior de Investigaciones Científicas - Universidad del País Vasco/Euskal Herriko Unibertsitatea (CSIC-UPV/EHU), Barrio Sarriena s/n, Leioa, Bizkaia 48940, Spain; Departamento de Bioquímica, Universidad del País Vasco, 48080 Bilbao, Spain; IKERBASQUE, Basque Foundation for Science, 48011 Bilbao, Spain.
Organizational Affiliation: