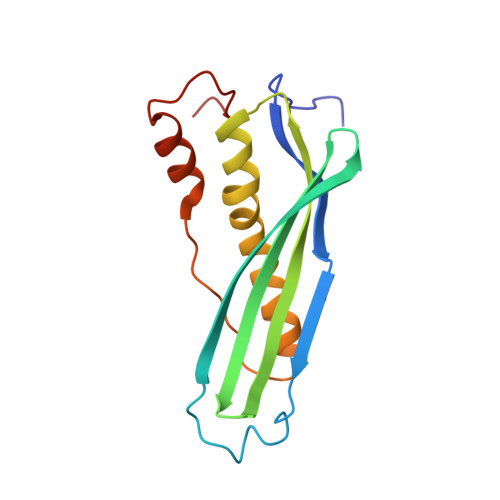
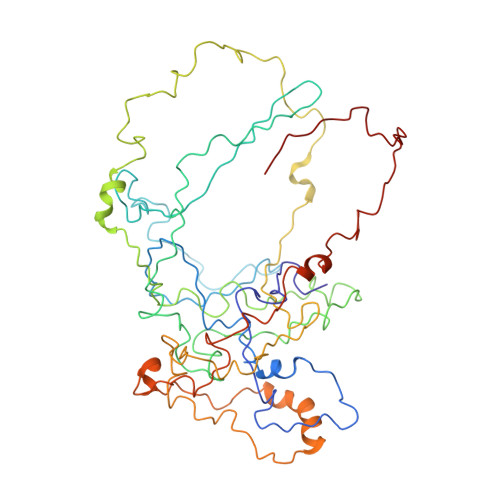
Structural basis for the antifolding activity of a molecular chaperone.
Huang, C., Rossi, P., Saio, T., Kalodimos, C.G.(2016) Nature 537: 202-206
- PubMed: 27501151 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature18965
- Primary Citation Related Structures:
5JTL, 5JTM, 5JTN, 5JTO, 5JTP, 5JTQ, 5JTR - PubMed Abstract:
Molecular chaperones act on non-native proteins in the cell to prevent their aggregation, premature folding or misfolding. Different chaperones often exert distinct effects, such as acceleration or delay of folding, on client proteins via mechanisms that are poorly understood. Here we report the solution structure of SecB, a chaperone that exhibits strong antifolding activity, in complex with alkaline phosphatase and maltose-binding protein captured in their unfolded states. SecB uses long hydrophobic grooves that run around its disk-like shape to recognize and bind to multiple hydrophobic segments across the length of non-native proteins. The multivalent binding mode results in proteins wrapping around SecB. This unique complex architecture alters the kinetics of protein binding to SecB and confers strong antifolding activity on the chaperone. The data show how the different architectures of chaperones result in distinct binding modes with non-native proteins that ultimately define the activity of the chaperone.
- Department of Biochemistry, Molecular Biology &Biophysics, University of Minnesota, Minneapolis, Minnesota 55455, USA.
Organizational Affiliation: