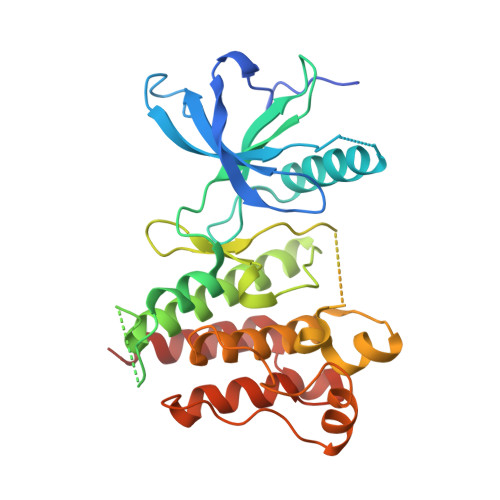
Crystal Structure of the FGFR4/LY2874455 Complex Reveals Insights into the Pan-FGFR Selectivity of LY2874455
Wu, D., Guo, M., Philips, M.A., Qu, L., Jiang, L., Li, J., Chen, X., Chen, Z., Chen, L., Chen, Y.(2016) PLoS One 11: e0162491-e0162491
- PubMed: 27618313 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0162491
- Primary Citation Related Structures:
5JKG - PubMed Abstract:
Aberrant FGFR4 signaling has been documented abundantly in various human cancers. The majority of FGFR inhibitors display significantly reduced potency toward FGFR4 compared to FGFR1-3. However, LY2874455 has similar inhibition potency for FGFR1-4 with IC50 less than 6.4 nM. To date, there is no published crystal structure of LY2874455 in complex with any kinase. To better understand the pan-FGFR selectivity of LY2874455, we have determined the crystal structure of the FGFR4 kinase domain bound to LY2874455 at a resolution of 2.35 Å. LY2874455, a type I inhibitor for FGFR4, binds to the ATP-binding pocket of FGFR4 in a DFG-in active conformation with three hydrogen bonds and a number of van der Waals contacts. After alignment of the kinase domain sequence of 4 FGFRs, and superposition of the ATP binding pocket of 4 FGFRs, our structural analyses reveal that the interactions of LY2874455 to FGFR4 are largely conserved in 4 FGFRs, explaining at least partly, the broad inhibitory activity of LY2874455 toward 4 FGFRs. Consequently, our studies reveal new insights into the pan-FGFR selectivity of LY2874455 and provide a structural basis for developing novel FGFR inhibitors that target FGFR1-4 broadly.
- Laboratory of Structural Biology, Key Laboratory of Cancer Proteomics of Chinese Ministry of Health, XiangYa Hospital, Central South University, Changsha, Hunan 410008, China.
Organizational Affiliation: