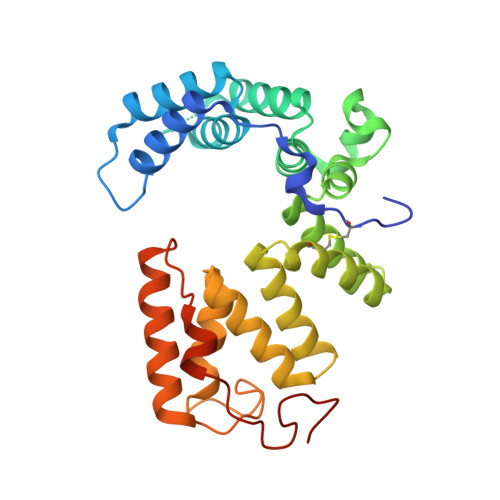
Structural analysis of Pseudomonas aeruginosa H3-T6SS immunity proteins
Yang, X.Y., Li, Z.Q., She, Z., Geng, Z., Xu, J.H., Gao, Z.Q., Dong, Y.H.(2016) FEBS Lett 590: 2787-2796
- PubMed: 27397502
- DOI: https://doi.org/10.1002/1873-3468.12291
- Primary Citation Related Structures:
5JJO, 5JKP - PubMed Abstract:
The Pseudomonas aeruginosa PldB protein is a transkingdom effector secreted by the Type VI Secretion System (T6SS). PA5088, PA5087, and PA5086 are three immunity proteins that can suppress the virulence of PldB. We report the crystal structures of PA5088 and PA5087 at 2.0 and 2.1 Å resolution, respectively. PA5088 and PA5087 both consist of several Sel1-like Repeats (SLRs) and form super-ring folds. Our structural analysis of these proteins revealed key differences among PA5088, PA5087, and their homologs. Our docking experiments have shed light on the putative interaction mechanism of their function as phospholipase D inhibitors.
- School of Life Sciences, University of Science and Technology of China, Hefei, China.
Organizational Affiliation: