Discovery of N-(3-(5-((3-acrylamido-4-(morpholine-4-carbonyl)phenyl)amino)-1-methyl-6-oxo-1,6-dihydropyridin-3-yl)-2-methylphenyl)-4-(tert-butyl)benzamide (CHMFL-BTK-01) as a highly selective irreversible Bruton's tyrosine kinase (BTK) inhibitor.
Liang, Q., Chen, Y., Yu, K., Chen, C., Zhang, S., Wang, A., Wang, W., Wu, H., Liu, X., Wang, B., Wang, L., Hu, Z., Wang, W., Ren, T., Zhang, S., Liu, Q., Yun, C.H., Liu, J.(2017) Eur J Med Chem 131: 107-125
- PubMed: 28315597 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2017.03.001
- Primary Citation Related Structures:
5J87 - PubMed Abstract:
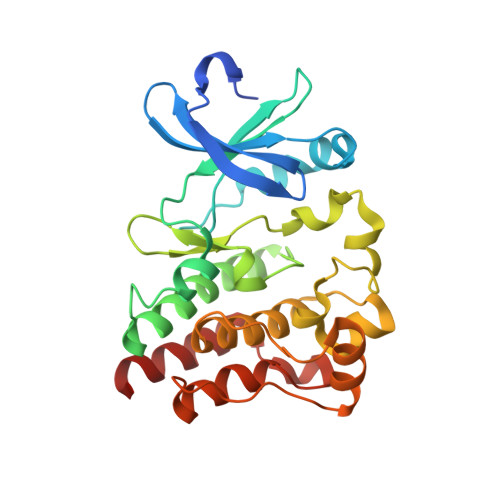
Currently there are several irreversible BTK inhibitors targeting Cys481 residue under preclinical or clinical development. However, most of these inhibitors also targeted other kinases such as BMX, JAK3, and EGFR that bear the highly similar active cysteine residues. Through a structure-based drug design approach, we discovered a highly potent (IC 50 : 7 nM) irreversible BTK inhibitor compound 9 (CHMFL-BTK-01), which displayed a high selectivity profile in KINOMEscan (S score (35) = 0.00) among 468 kinases/mutants at the concentration of 1 μM. Compound 9 completely abolished BMX, JAK3 and EGFR's activity. Both X-ray crystal structure and cysteine-serine mutation mediated rescue experiment confirmed 9's irreversible binding mode. 9 also potently inhibited BTK Y223 auto-phosphorylation (EC 50 : <30 nM), arrested cell cycle in G0/G1 phase and induced apoptosis in U2932 and Pfeiffer cells. We believe these features would make 9 a good pharmacological tool to study the BTK related pathology.
- Department of Chemistry, University of Science and Technology of China, Hefei, Anhui 230036, PR China; High Magnetic Field Laboratory, Chinese Academy of Sciences, Mailbox 1110, 350 Shushanhu Road, Hefei, Anhui 230031, PR China.
Organizational Affiliation: