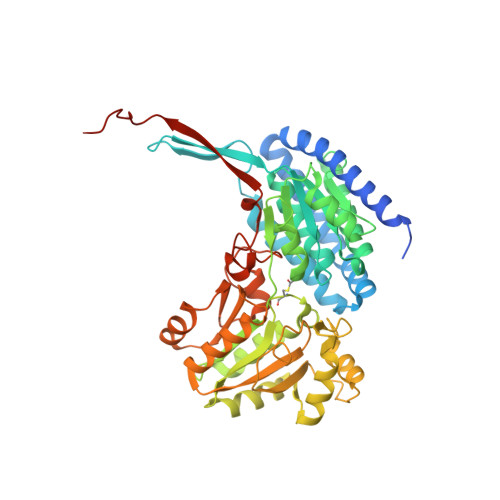
Structure of an acetylating aldehyde dehydrogenase from the thermophilic ethanologen Geobacillus thermoglucosidasius.
Extance, J., Danson, M.J., Crennell, S.J.(2016) Protein Sci 25: 2045-2053
- PubMed: 27571338 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3027
- Primary Citation Related Structures:
5J78, 5J7I - PubMed Abstract:
Acetylating aldehyde dehydrogenases (AcAldDH) catalyse the acetylation of Coenzyme-A (CoA), or in reverse generate acetaldehyde from Acetyl-CoA using NADH as a co-factor. This article reports the expression, purification, enzyme assay, and X-ray crystal structures of an AcAldDH from Geobacillus thermoglucosidasius (GtAcAldDH) to 2.1Å and in complex with CoA and NAD + to 4.0Å. In the structure, the AcAldDH forms a close-knit dimer, similar to that seen in other Alcohol Dehydrogenase (ADH) structures. In GtAcAldDH, these dimers associate via their N-termini to form weakly interacting tetramers. This mode of tetrameric association is also seen in an unpublished AcAldDH deposited in the PDB, but is in contrast to all other ADH structures, (including the one other published AcAldDH found in a bacterial microcompartment), in which the dimers bury a large surface area including the C-termini. This novel mode of association sequesters the active sites and potentially reactive acyl-enzyme intermediates in the center of the tetramer. In other respects, the structure is very similar to the other AcAldDH, binding the cofactors in a corresponding fashion. This similarity enabled the identification of a shortened substrate cavity in G. thermoglucosidasius AcAldDH, explaining the limitations on the length of substrate accepted by the enzyme.
- Centre for Extremophile Research, Department of Biology and Biochemistry, University of Bath, Bath, BA2 7AY, England.
Organizational Affiliation: