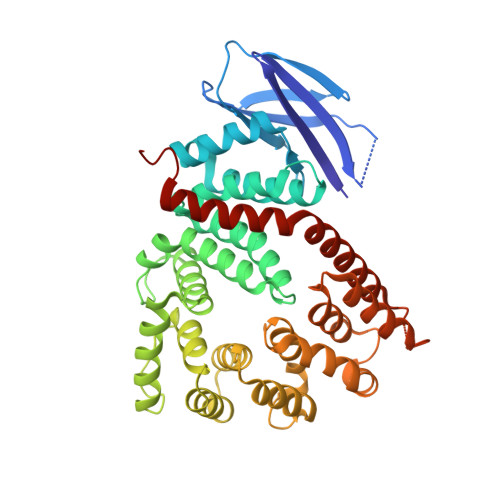
Crystal structure of a polypeptide's C-terminus in complex with the regulatory domain of ER aminopeptidase 1.
Sui, L., Gandhi, A., Guo, H.C.(2016) Mol Immunol 80: 41-49
- PubMed: 27825049 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molimm.2016.10.012
- Primary Citation Related Structures:
5J5E - PubMed Abstract:
Endoplasmic reticulum aminopeptidase 1 (ERAP1) is involved in the final processing of peptide precursors to generate the N-termini of MHC class I-restricted epitopes. ERAP1 thus influences immunodominance and cytotoxic immune responses by controlling the peptide repertoire available for cell surface presentation by MHC molecules. To enable this critical role in antigen processing, ERAP1 trims peptides by a unique molecular ruler mechanism that turns on/off hydrolysis activity in a peptide-length and -sequence dependent manner. Thus unlike other aminopeptidases, ERAP1 could recognize both the N- and C-termini of peptides in order to read the substrate's length. To exemplify and validate this molecular ruler mechanism, we have carried out crystallographic studies on molecular recognition of antigenic peptide's C-terminus by ERAP1. In this report, we have determined a 2.8Å-resolution crystal structure of an intermolecular complex between the ERAP1 regulatory domain and a natural epitope's C-terminus displayed in a fusion protein. It reveals the structural details of peptide's C-termini recognition by ERAP1. ERAP1 uses specificity pockets on the regulatory domain to bind the peptide's carboxyl end and side chain of the C-terminal anchoring residue. At the same time, flexibility in length and sequence at the middle of peptides is accommodated by a kink with minimal interactions with ERAP1.
- Department of Biological Sciences, University of Massachusetts Lowell, 1 University Avenue, Lowell, MA 01854, USA.
Organizational Affiliation: