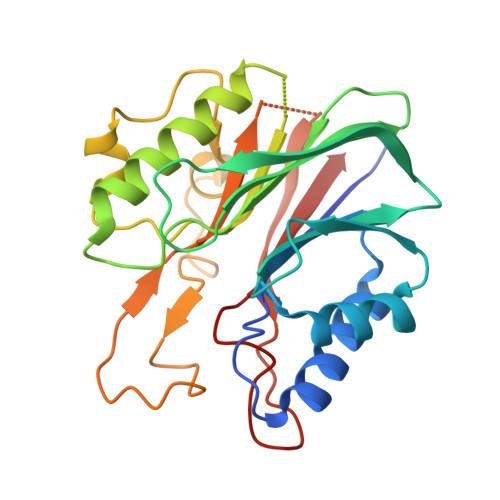
Mode of action of DNA-competitive small molecule inhibitors of tyrosyl DNA phosphodiesterase 2.
Hornyak, P., Askwith, T., Walker, S., Komulainen, E., Paradowski, M., Pennicott, L.E., Bartlett, E.J., Brissett, N.C., Raoof, A., Watson, M., Jordan, A.M., Ogilvie, D.J., Ward, S.E., Atack, J.R., Pearl, L.H., Caldecott, K.W., Oliver, A.W.(2016) Biochem J 473: 1869-1879
- PubMed: 27099339 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20160180
- Primary Citation Related Structures:
5J3P, 5J3S, 5J3Z, 5J42 - PubMed Abstract:
Tyrosyl-DNA phosphodiesterase 2 (TDP2) is a 5'-tyrosyl DNA phosphodiesterase important for the repair of DNA adducts generated by non-productive (abortive) activity of topoisomerase II (TOP2). TDP2 facilitates therapeutic resistance to topoisomerase poisons, which are widely used in the treatment of a range of cancer types. Consequently, TDP2 is an interesting target for the development of small molecule inhibitors that could restore sensitivity to topoisomerase-directed therapies. Previous studies identified a class of deazaflavin-based molecules that showed inhibitory activity against TDP2 at therapeutically useful concentrations, but their mode of action was uncertain. We have confirmed that the deazaflavin series inhibits TDP2 enzyme activity in a fluorescence-based assay, suitable for high-throughput screen (HTS)-screening. We have gone on to determine crystal structures of these compounds bound to a 'humanized' form of murine TDP2. The structures reveal their novel mode of action as competitive ligands for the binding site of an incoming DNA substrate, and point the way to generating novel and potent inhibitors of TDP2.
- Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer BN1 9RQ, U.K. Cancer Research UK DNA Repair Enzymes Group, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer BN1 9RQ, U.K.
Organizational Affiliation: