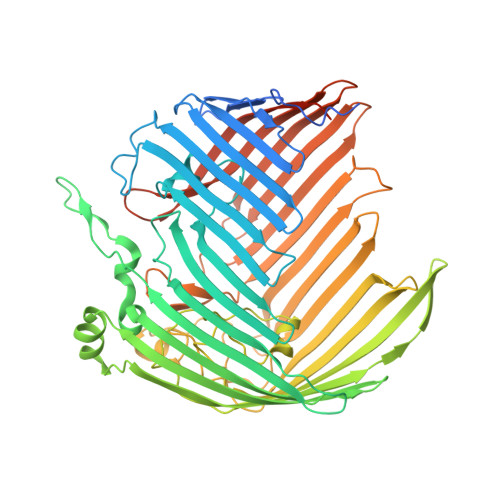
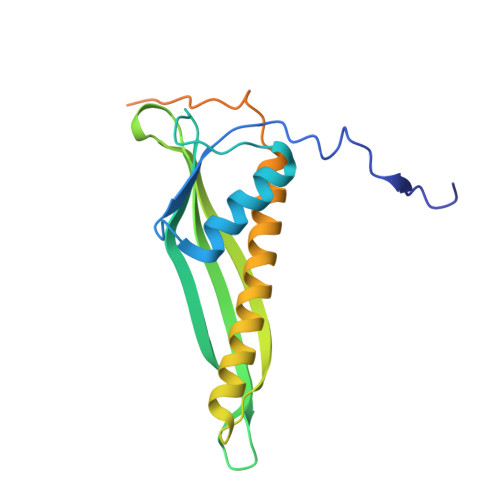
Structural and Functional Characterization of the LPS Transporter LptDE from Gram-Negative Pathogens.
Botos, I., Majdalani, N., Mayclin, S.J., McCarthy, J.G., Lundquist, K., Wojtowicz, D., Barnard, T.J., Gumbart, J.C., Buchanan, S.K.(2016) Structure 24: 965-976
- PubMed: 27161977 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.03.026
- Primary Citation Related Structures:
5IV8, 5IV9, 5IVA, 5IXM - PubMed Abstract:
Incorporation of lipopolysaccharide (LPS) into the outer membrane of Gram-negative bacteria is essential for viability, and is accomplished by a two-protein complex called LptDE. We solved crystal structures of the core LptDE complexes from Yersinia pestis, Klebsiella pneumoniae, Pseudomonas aeruginosa, and a full-length structure of the K. pneumoniae LptDE complex. Our structures adopt the same plug and 26-strand β-barrel architecture found recently for the Shigella flexneri and Salmonella typhimurium LptDE structures, illustrating a conserved fold across the family. A comparison of the only two full-length structures, SfLptDE and our KpLptDE, reveals a 21° rotation of the LptD N-terminal domain that may impart flexibility on the trans-envelope LptCAD scaffold. Utilizing mutagenesis coupled to an in vivo functional assay and molecular dynamics simulations, we demonstrate the critical role of Pro231 and Pro246 in the function of the LptD lateral gate that allows partitioning of LPS into the outer membrane.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: