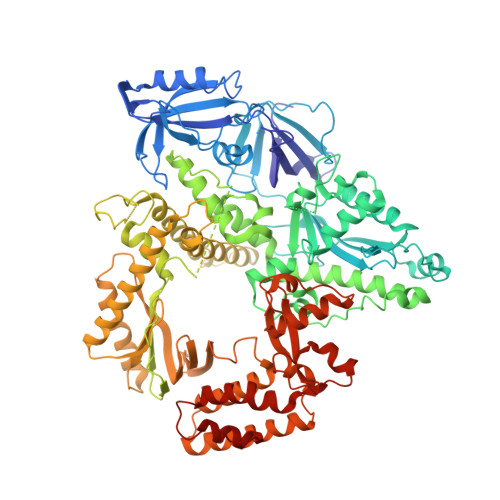
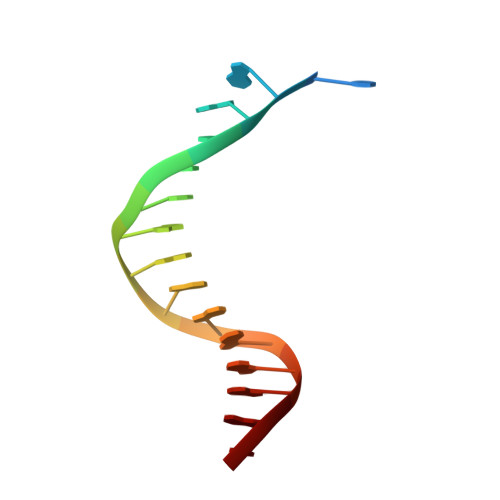
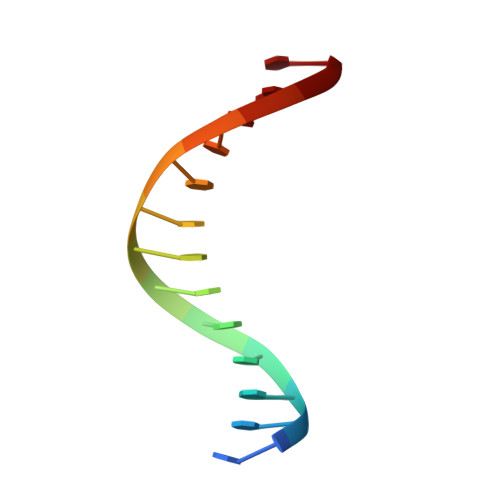
Human DNA polymerase alpha in binary complex with a DNA:DNA template-primer.
Coloma, J., Johnson, R.E., Prakash, L., Prakash, S., Aggarwal, A.K.(2016) Sci Rep 6: 23784-23784
- PubMed: 27032819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep23784
- Primary Citation Related Structures:
5IUD - PubMed Abstract:
The Polα/primase complex assembles the short RNA-DNA fragments for priming of lagging and leading strand DNA replication in eukaryotes. As such, the Polα polymerase subunit encounters two types of substrates during primer synthesis: an RNA:DNA helix and a DNA:DNA helix. The engagement of the polymerase subunit with the DNA:DNA helix has been suggested as the of basis for primer termination in eukaryotes. However, there is no structural information on how the Polα polymerase subunit actually engages with a DNA:DNA helix during primer synthesis. We present here the first crystal structure of human Polα polymerase subunit in complex with a DNA:DNA helix. Unexpectedly, we find that portion of the DNA:DNA helix in contact with the polymerase is not in a B-form but in a hybrid A-B form. Almost all of the contacts observed previously with an RNA primer are preserved with a DNA primer--with the same set of polymerase residues tracking the sugar-phosphate backbone of the DNA or RNA primer. Thus, rather than loss of specific contacts, the free energy cost of distorting DNA from B- to hybrid A-B form may augur the termination of primer synthesis in eukaryotes.
- Department of Structural &Chemical Biology, Mount Sinai School of Medicine, Box 1677, 1425 Madison Avenue, New York, NY 10029, USA.
Organizational Affiliation: