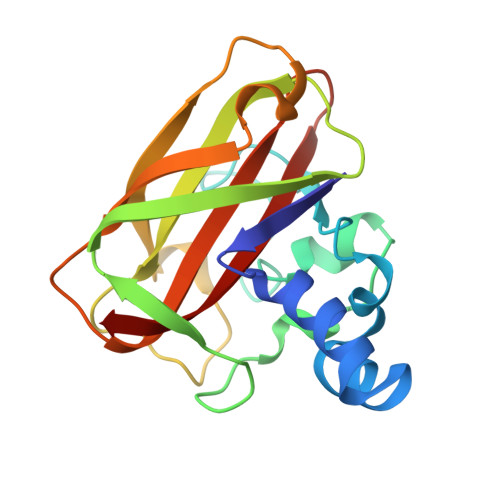
Activity, stability and 3-D structure of the Cu(ii) form of a chitin-active lytic polysaccharide monooxygenase from Bacillus amyloliquefaciens.
Gregory, R.C., Hemsworth, G.R., Turkenburg, J.P., Hart, S.J., Walton, P.H., Davies, G.J.(2016) Dalton Trans 45: 16904-16912
- PubMed: 27722375 Search on PubMed
- DOI: https://doi.org/10.1039/c6dt02793h
- Primary Citation Related Structures:
5IJU - PubMed Abstract:
The enzymatic deconstruction of recalcitrant polysaccharide biomass is central to the conversion of these substrates for societal benefit, such as in biofuels. Traditional models for enzyme-catalysed polysaccharide degradation involved the synergistic action of endo-, exo- and processive glycoside hydrolases working in concert to hydrolyse the substrate. More recently this model has been succeeded by one featuring a newly discovered class of mononuclear copper enzymes: lytic polysaccharide monooxygenases (LPMOs; classified as Auxiliary Activity (AA) enzymes in the CAZy classification). In 2013, the structure of an LPMO from Bacillus amyloliquefaciens, BaAA10, was solved with the Cu centre photoreduced to Cu(i) in the X-ray beam. Here we present the catalytic activity of BaAA10. We show that it is a chitin-active LPMO, active on both α and β chitin, with the Cu(ii) binding with low nM K D , and the substrate greatly increasing the thermal stability of the enzyme. A spiral data collection strategy has been used to facilitate access to the previously unobservable Cu(ii) state of the active centre, revealing a coordination geometry around the copper which is distorted from axial symmetry, consistent with the previous findings from EPR spectroscopy.
- Structural Biology Laboratory, University of York, Heslington, York, YO10 5DD, UK. gideon.davies@york.ac.uk.
Organizational Affiliation: