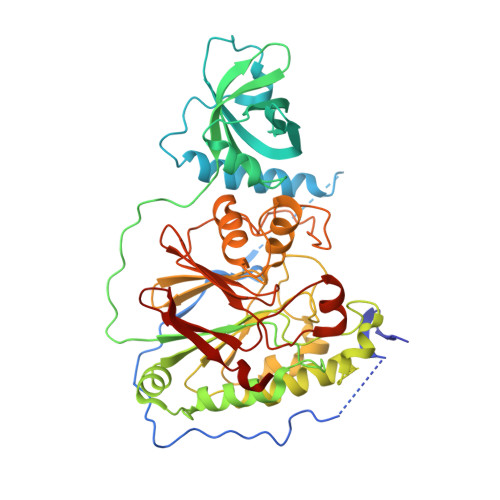
Shared active site architecture between archaeal PolD and multi-subunit RNA polymerases revealed by X-ray crystallography.
Sauguet, L., Raia, P., Henneke, G., Delarue, M.(2016) Nat Commun 7: 12227-12227
- PubMed: 27548043 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms12227
- Primary Citation Related Structures:
5IHE, 5IJL - PubMed Abstract:
Archaeal replicative DNA polymerase D (PolD) constitute an atypical class of DNA polymerases made of a proofreading exonuclease subunit (DP1) and a larger polymerase catalytic subunit (DP2), both with unknown structures. We have determined the crystal structures of Pyrococcus abyssi DP1 and DP2 at 2.5 and 2.2 Å resolution, respectively, revealing a catalytic core strikingly different from all other known DNA polymerases (DNAPs). Rather, the PolD DP2 catalytic core has the same 'double-psi β-barrel' architecture seen in the RNA polymerase (RNAP) superfamily, which includes multi-subunit transcriptases of all domains of life, homodimeric RNA-silencing pathway RNAPs and atypical viral RNAPs. This finding bridges together, in non-viral world, DNA transcription and DNA replication within the same protein superfamily. This study documents further the complex evolutionary history of the DNA replication apparatus in different domains of life and proposes a classification of all extant DNAPs.
- Unit of Structural Dynamics of Macromolecules, Pasteur Institute and CNRS UMR 3528, 75015 Paris, France.
Organizational Affiliation: