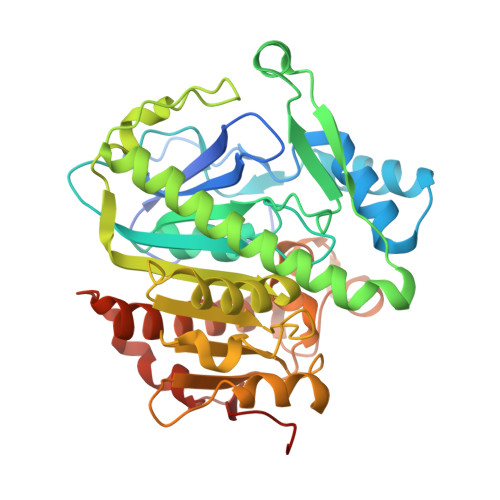
Structural and Thermodynamic Characterization of Protein-Ligand Interactions Formed between Lipoprotein-Associated Phospholipase A2 and Inhibitors
Liu, Q.F., Chen, X.D., Chen, W.Y., Yuan, X.J., Su, H.X., Shen, J.H., Xu, Y.C.(2016) J Med Chem 59: 5115-5120
- PubMed: 27078579 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b00282
- Primary Citation Related Structures:
5I8P, 5I9I - PubMed Abstract:
Lipoprotein-associated phospholipase A2 (Lp-PLA2) represents a promising therapeutic target for atherosclerosis and Alzheimer's disease. Here we reported the first crystal structures of Lp-PLA2 bound with reversible inhibitors and the thermodynamic characterization of complexes. High rigidity of Lp-PLA2 structure and similar binding modes of inhibitors with completely different scaffolds are revealed. It not only provides the molecular basis for inhibitory activity but also sheds light on the essential features of Lp-PLA2 recognition with reversible inhibitors.
- School of Life Science and Technology, ShanghaiTech University , Shanghai 200031, China.
Organizational Affiliation: