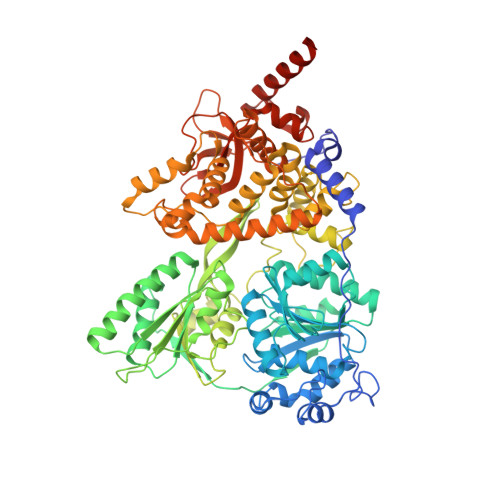
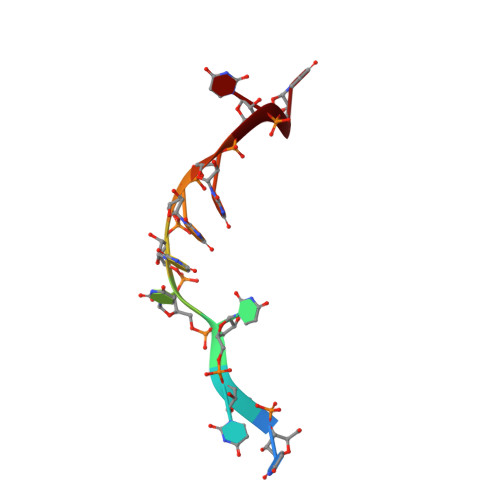
Structure of the DEAH/RHA ATPase Prp43p bound to RNA implicates a pair of hairpins and motif Va in translocation along RNA.
He, Y., Staley, J.P., Andersen, G.R., Nielsen, K.H.(2017) RNA 23: 1110-1124
- PubMed: 28416566 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.060954.117
- Primary Citation Related Structures:
5I8Q - PubMed Abstract:
Three families of nucleic acid-dependent ATPases (DEAH/RHA, Ski2-like, and NS3/NPH-II), termed the DExH ATPases, are thought to execute myriad functions by processive, ATP-dependent, 3' to 5' translocation along single-stranded nucleic acid. While the mechanism of translocation of the viral NS3/NPH-II family has been studied extensively, it has not been clear if or how the principles that have emerged for this family extend to the other two families. Here we report the crystal structure of the yeast DEAH/RHA family ATPase Prp43p, which functions in splicing and ribosome biogenesis, in complex with poly-uracil and a nonhydrolyzable ATP analog. The structure reveals a conserved DEAH/RHA-specific variation of motif Ib within the RecA1 domain of the catalytic core, in which the motif elongates as a β-hairpin that bookends the 3' end of a central RNA stack, a function that in the viral and Ski-2 families is performed by an auxiliary domain. Supporting a fundamental role in translocation, mutations in this hairpin abolished helicase activity without affecting RNA binding or ATPase activity. While the structure reveals differences with viral ATPases in the RecA1 domain, our structure demonstrates striking similarities with viral ATPases in the RecA2 domain of the catalytic core, including both a prominent β-hairpin that bookends the 5' end of the RNA stack and a dynamic motif Va that is implicated in mediating translocation. Our crystal structure, genetic, and biochemical experiments, as well as comparisons with other DExH ATPases, support a generalized mechanism for the DExH class of helicases involving a pair of bookends that inchworm along RNA.
- Department of Molecular Biology and Genetics, Aarhus University, DK8000 Aarhus C, Denmark.
Organizational Affiliation: