Bacterial Chaperone Domain Insertions Convert Human FKBP12 into an Excellent Protein-Folding Catalyst-A Structural and Functional Analysis.
Zoldak, G., Knappe, T.A., Geitner, A.J., Scholz, C., Dobbek, H., Schmid, F.X., Jakob, R.P.(2024) Molecules 29
- PubMed: 38611720 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules29071440
- Primary Citation Related Structures:
5I7P, 5I7Q - PubMed Abstract:
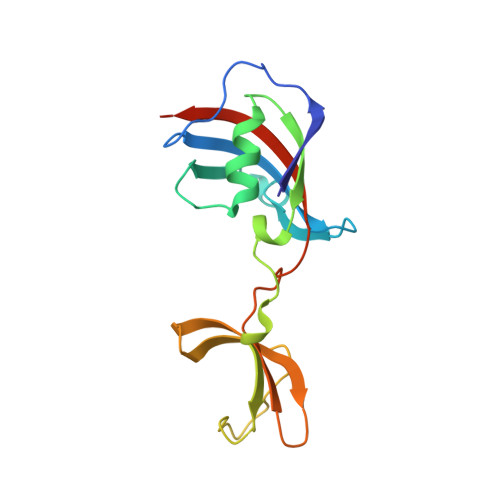
Many folding enzymes use separate domains for the binding of substrate proteins and for the catalysis of slow folding reactions such as prolyl isomerization. FKBP12 is a small prolyl isomerase without a chaperone domain. Its folding activity is low, but it could be increased by inserting the chaperone domain from the homolog SlyD of E. coli near the prolyl isomerase active site. We inserted two other chaperone domains into human FKBP12: the chaperone domain of SlpA from E. coli , and the chaperone domain of SlyD from Thermococcus sp. Both stabilized FKBP12 and greatly increased its folding activity. The insertion of these chaperone domains had no influence on the FKBP12 and the chaperone domain structure, as revealed by two crystal structures of the chimeric proteins. The relative domain orientations differ in the two crystal structures, presumably representing snapshots of a more open and a more closed conformation. Together with crystal structures from SlyD-like proteins, they suggest a path for how substrate proteins might be transferred from the chaperone domain to the prolyl isomerase domain.
- Center for Interdisciplinary Biosciences, Technology and Innovation Park, Pavol Jozef Šafárik University in Košice, 040 11 Kosice, Slovakia.
Organizational Affiliation: