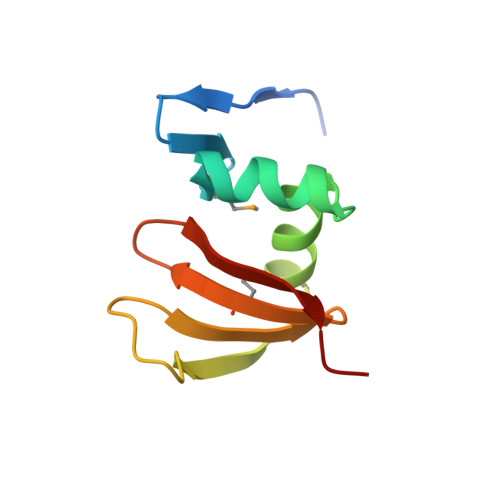
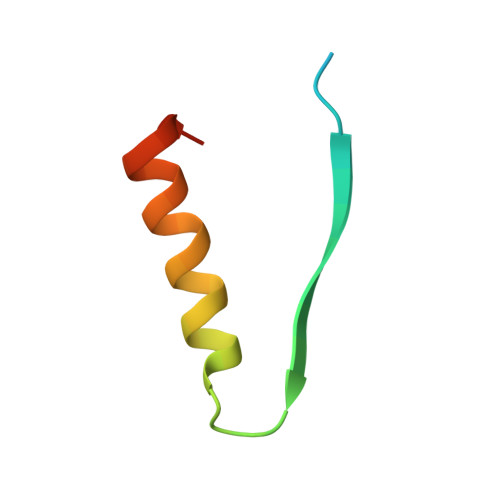
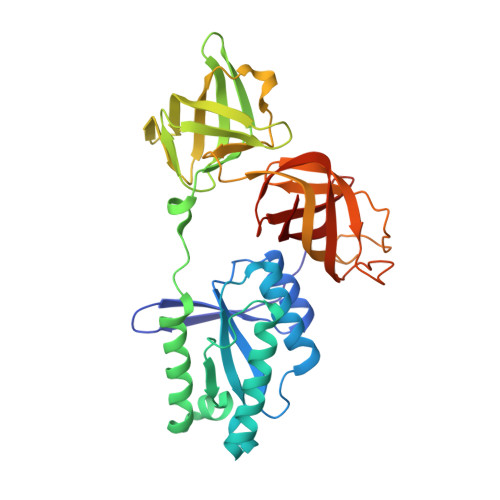
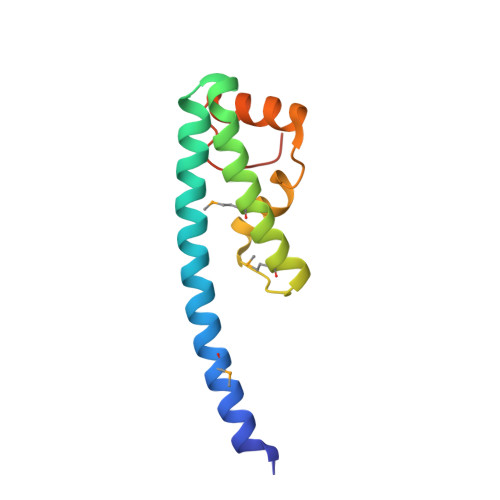
Structure of a novel antibacterial toxin that exploits elongation factor Tu to cleave specific transfer RNAs.
Michalska, K., Gucinski, G.C., Garza-Sanchez, F., Johnson, P.M., Stols, L.M., Eschenfeldt, W.H., Babnigg, G., Low, D.A., Goulding, C.W., Joachimiak, A., Hayes, C.S.(2017) Nucleic Acids Res 45: 10306-10320
- PubMed: 28973472 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkx700
- Primary Citation Related Structures:
5I4Q, 5I4R - PubMed Abstract:
Contact-dependent growth inhibition (CDI) is a mechanism of inter-cellular competition in which Gram-negative bacteria exchange polymorphic toxins using type V secretion systems. Here, we present structures of the CDI toxin from Escherichia coli NC101 in ternary complex with its cognate immunity protein and elongation factor Tu (EF-Tu). The toxin binds exclusively to domain 2 of EF-Tu, partially overlapping the site that interacts with the 3'-end of aminoacyl-tRNA (aa-tRNA). The toxin exerts a unique ribonuclease activity that cleaves the single-stranded 3'-end from tRNAs that contain guanine discriminator nucleotides. EF-Tu is required to support this tRNase activity in vitro, suggesting the toxin specifically cleaves substrate in the context of GTP·EF-Tu·aa-tRNA complexes. However, superimposition of the toxin domain onto previously solved GTP·EF-Tu·aa-tRNA structures reveals potential steric clashes with both aa-tRNA and the switch I region of EF-Tu. Further, the toxin induces conformational changes in EF-Tu, displacing a β-hairpin loop that forms a critical salt-bridge contact with the 3'-terminal adenylate of aa-tRNA. Together, these observations suggest that the toxin remodels GTP·EF-Tu·aa-tRNA complexes to free the 3'-end of aa-tRNA for entry into the nuclease active site.
- Midwest Center for Structural Genomics, Biosciences Division, Argonne National Laboratory, Argonne, IL 60439, USA.
Organizational Affiliation: