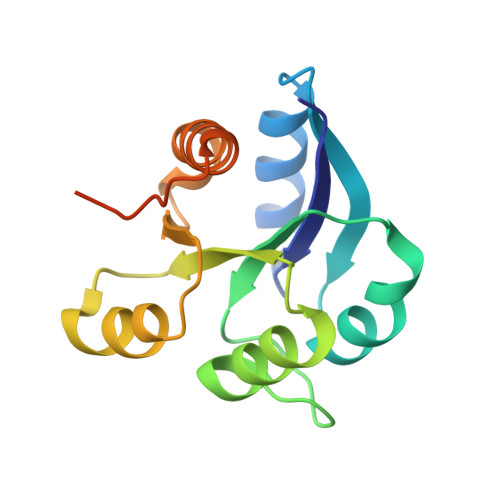
Crystal structure of nonphosphorylated receiver domain of the stress response regulator RcsB from Escherichia coli.
Filippova, E.V., Wawrzak, Z., Ruan, J., Pshenychnyi, S., Schultz, R.M., Wolfe, A.J., Anderson, W.F.(2016) Protein Sci 25: 2216-2224
- PubMed: 27670836
- DOI: https://doi.org/10.1002/pro.3050
- Primary Citation Related Structures:
5I4C - PubMed Abstract:
RcsB, the transcription-associated response regulator of the Rcs phosphorelay two-component signal transduction system, activates cell stress responses associated with desiccation, cell wall biosynthesis, cell division, virulence, biofilm formation, and antibiotic resistance in enteric bacterial pathogens. RcsB belongs to the FixJ/NarL family of transcriptional regulators, which are characterized by a highly conserved C-terminal DNA-binding domain. The N-terminal domain of RcsB belongs to the family of two-component receiver domains. This receiver domain contains the phosphoacceptor site and participates in RcsB dimer formation; it also contributes to dimer formation with other transcription factor partners. Here, we describe the crystal structure of the Escherichia coli RcsB receiver domain in its nonphosphorylated state. The structure reveals important molecular details of phosphorylation-independent dimerization of RcsB and has implication for the formation of heterodimers.
- Department of Biochemistry and Molecular Genetics, Center for Structural Genomics of Infectious Diseases, Northwestern University Feinberg School of Medicine, Chicago, Illinois, 60611.
Organizational Affiliation: