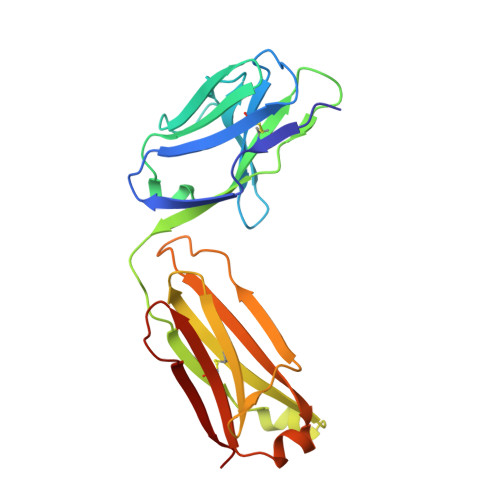
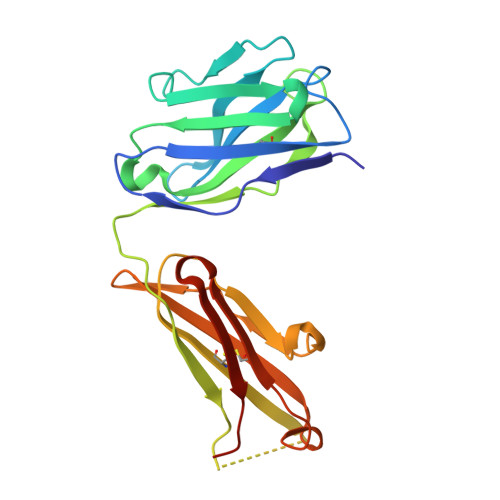
De Novo Sequencing and Resurrection of a Human Astrovirus-Neutralizing Antibody.
Bogdanoff, W.A., Morgenstern, D., Bern, M., Ueberheide, B.M., Sanchez-Fauquier, A., DuBois, R.M.(2016) ACS Infect Dis 2: 313-321
- PubMed: 27213181 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsinfecdis.6b00026
- Primary Citation Related Structures:
5I30 - PubMed Abstract:
Monoclonal antibody (mAb) therapeutics targeting cancer, autoimmune diseases, inflammatory diseases, and infectious diseases are growing exponentially. Although numerous panels of mAbs targeting infectious disease agents have been developed, their progression into clinically useful mAbs is often hindered by the lack of sequence information and/or loss of hybridoma cells that produce them. Here we combine the power of crystallography and mass spectrometry to determine the amino acid sequence and glycosylation modification of the Fab fragment of a potent human astrovirus-neutralizing mAb. We used this information to engineer a recombinant antibody single-chain variable fragment that has the same specificity as the parent monoclonal antibody to bind to the astrovirus capsid protein. This antibody can now potentially be developed as a therapeutic and diagnostic agent.
- Department of Biomolecular Engineering, University of California Santa Cruz , 1156 High Street, Santa Cruz, California 95064, United States.
Organizational Affiliation: