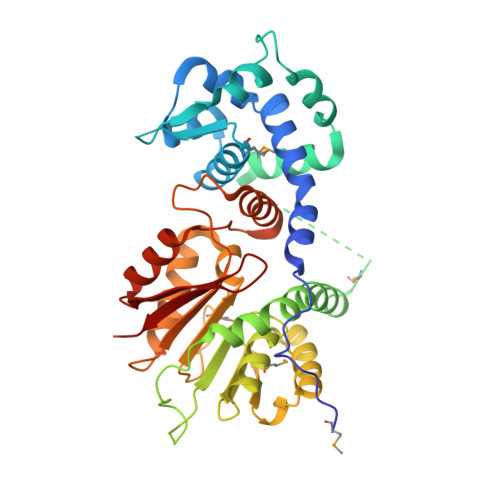
Crystal structure of O-methyltransferase family 2 protein Plim_1147 from Planctomyces limnophilus DSM 3776 complex with Apigenin.
Chang, C., Duke, N., Bigelow, L., Bearden, J., Joachimiak, A., Midwest Center for Structural Genomics (MCSG)To be published.