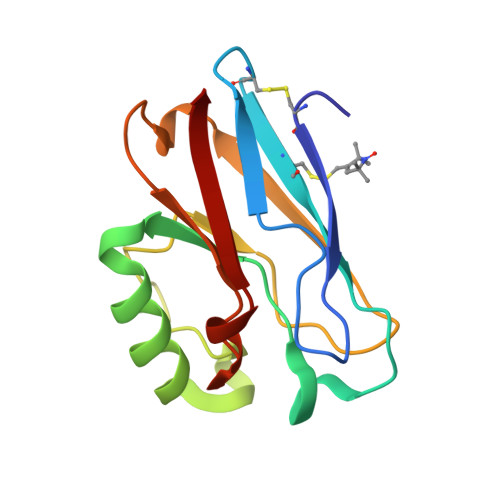
Determination of nitroxide spin label conformations via PELDOR and X-ray crystallography.
Abdullin, D., Hagelueken, G., Schiemann, O.(2016) Phys Chem Chem Phys 18: 10428-10437
- PubMed: 27029516 Search on PubMed
- DOI: https://doi.org/10.1039/c6cp01307d
- Primary Citation Related Structures:
5I26, 5I28 - PubMed Abstract:
Pulsed electron-electron double resonance (PELDOR or DEER) in combination with site-directed spin labelling has emerged as an important method for measuring nanometer distance constraints that are used to obtain coarse-grained structures of biomolecules or to follow their conformational changes. Translating measured spin-spin distances between spin labels into structural information requires taking the conformational flexibility of spin label side chains into account. Here, we present an analysis of orientation selective PELDOR data recorded on six singly MTSSL-labelled azurin mutants. The analysis yielded conformational MTSSL ensembles, which are considerably narrower than those predicted using in silico spin labeling methods but match well with spin label conformations found in the corresponding crystal structures. The possible reasons and consequences for predicting spin label conformers in the fold of biomolecules are discussed.
- Institute of Physical and Theoretical Chemistry, University of Bonn, Wegelerstr. 12, 53115 Bonn, Germany. schiemann@pc.uni-bonn.de.
Organizational Affiliation: