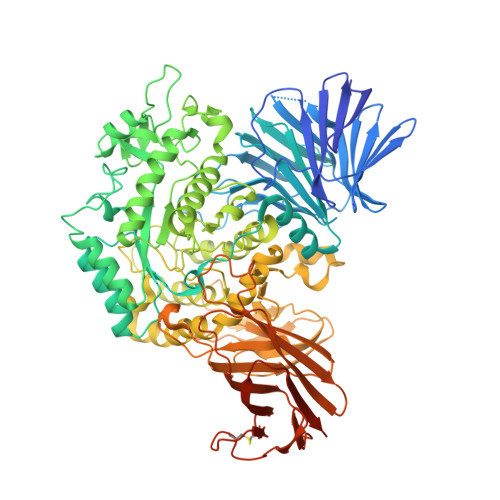
Detection of Active Mammalian GH31 alpha-Glucosidases in Health and Disease Using In-Class, Broad-Spectrum Activity-Based Probes.
Jiang, J., Kuo, C.L., Wu, L., Franke, C., Kallemeijn, W.W., Florea, B.I., van Meel, E., van der Marel, G.A., Codee, J.D., Boot, R.G., Davies, G.J., Overkleeft, H.S., Aerts, J.M.(2016) ACS Cent Sci 2: 351-358
- PubMed: 27280170 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acscentsci.6b00057
- Primary Citation Related Structures:
5I23, 5I24 - PubMed Abstract:
The development of small molecule activity-based probes (ABPs) is an evolving and powerful area of chemistry. There is a major need for synthetically accessible and specific ABPs to advance our understanding of enzymes in health and disease. α-Glucosidases are involved in diverse physiological processes including carbohydrate assimilation in the gastrointestinal tract, glycoprotein processing in the endoplasmic reticulum (ER), and intralysosomal glycogen catabolism. Inherited deficiency of the lysosomal acid α-glucosidase (GAA) causes the lysosomal glycogen storage disorder, Pompe disease. Here, we design a synthetic route for fluorescent and biotin-modified ABPs for in vitro and in situ monitoring of α-glucosidases. We show, through mass spectrometry, gel electrophoresis, and X-ray crystallography, that α-glucopyranose configured cyclophellitol aziridines label distinct retaining α-glucosidases including GAA and ER α-glucosidase II, and that this labeling can be tuned by pH. We illustrate a direct diagnostic application in Pompe disease patient cells, and discuss how the probes may be further exploited for diverse applications.
- Department of Bio-organic Synthesis, Leiden Institute of Chemistry, Leiden University , Einsteinweg 55, 2333 CC Leiden, The Netherlands.
Organizational Affiliation: