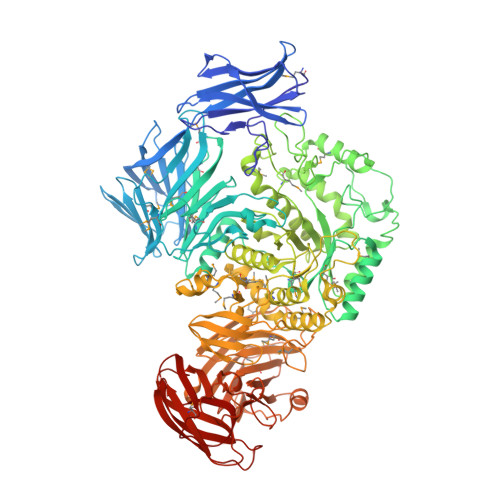
Transferase Versus Hydrolase: The Role of Conformational Flexibility in Reaction Specificity.
Light, S.H., Cahoon, L.A., Mahasenan, K.V., Lee, M., Boggess, B., Halavaty, A.S., Mobashery, S., Freitag, N.E., Anderson, W.F.(2017) Structure 25: 295-304
- PubMed: 28089449 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.12.007
- Primary Citation Related Structures:
5HOP, 5HPO, 5HXM, 5I0D, 5I0E, 5I0F, 5I0G - PubMed Abstract:
Active in the aqueous cellular environment where a massive excess of water is perpetually present, enzymes that catalyze the transfer of an electrophile to a non-water nucleophile (transferases) require specific strategies to inhibit mechanistically related hydrolysis reactions. To identify principles that confer transferase versus hydrolase reaction specificity, we exploited two enzymes that use highly similar catalytic apparatuses to catalyze the transglycosylation (a transferase reaction) or hydrolysis of α-1,3-glucan linkages in the cyclic tetrasaccharide cycloalternan (CA). We show that substrate binding to non-catalytic domains and a conformationally stable active site promote CA transglycosylation, whereas a distinct pattern of active site conformational change is associated with CA hydrolysis. These findings defy the classic view of induced-fit conformational change and illustrate a mechanism by which a stable hydrophobic binding site can favor transferase activity and disfavor hydrolysis. Application of these principles could facilitate the rational reengineering of transferases with desired catalytic properties.
- Department of Biochemistry and Molecular Genetics, Center for Structural Genomics of Infectious Diseases, Feinberg School of Medicine, Northwestern University, Chicago, IL 60611, USA.
Organizational Affiliation: