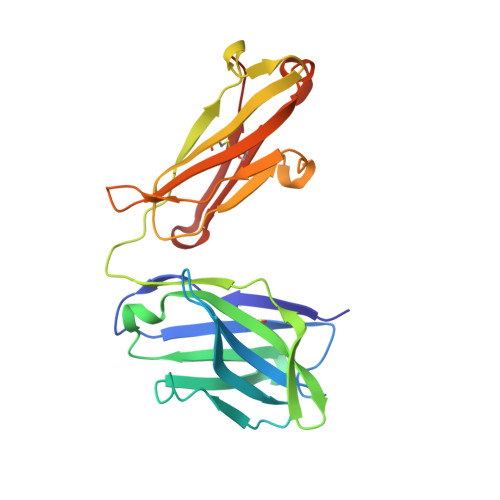
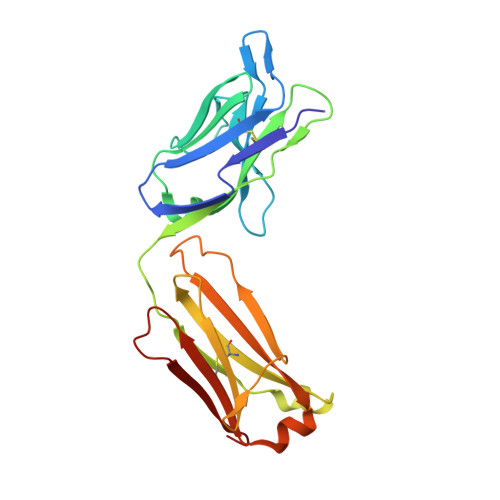
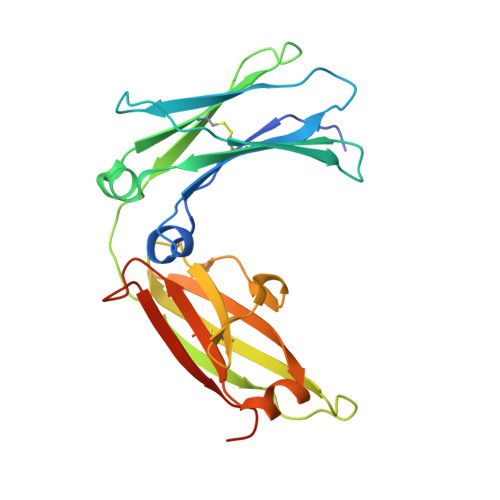
Structural basis of omalizumab therapy and omalizumab-mediated IgE exchange.
Pennington, L.F., Tarchevskaya, S., Brigger, D., Sathiyamoorthy, K., Graham, M.T., Nadeau, K.C., Eggel, A., Jardetzky, T.S.(2016) Nat Commun 7: 11610-11610
- PubMed: 27194387 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms11610
- Primary Citation Related Structures:
5HYS - PubMed Abstract:
Omalizumab is a widely used therapeutic anti-IgE antibody. Here we report the crystal structure of the omalizumab-Fab in complex with an IgE-Fc fragment. This structure reveals the mechanism of omalizumab-mediated inhibition of IgE interactions with both high- and low-affinity IgE receptors, and explains why omalizumab selectively binds free IgE. The structure of the complex also provides mechanistic insight into a class of disruptive IgE inhibitors that accelerate the dissociation of the high-affinity IgE receptor from IgE. We use this structural data to generate a mutant IgE-Fc fragment that is resistant to omalizumab binding. Treatment with this omalizumab-resistant IgE-Fc fragment, in combination with omalizumab, promotes the exchange of cell-bound full-length IgE with omalizumab-resistant IgE-Fc fragments on human basophils. This combination treatment also blocks basophil activation more efficiently than either agent alone, providing a novel approach to probe regulatory mechanisms underlying IgE hypersensitivity with implications for therapeutic interventions.
- Department of Structural Biology, Stanford University School of Medicine, Stanford, California 94305, USA.
Organizational Affiliation: