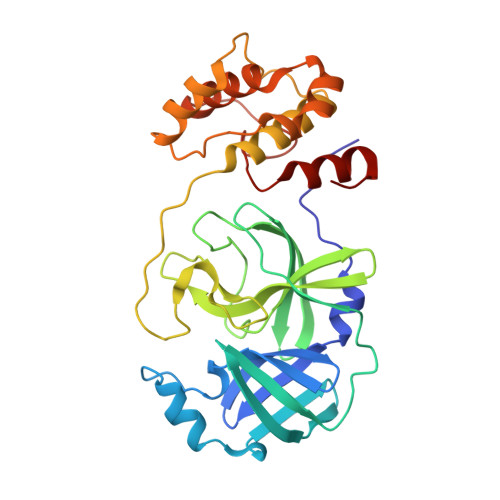
X-Ray Structure and Inhibition of 3C-like Protease from Porcine Epidemic Diarrhea Virus.
St John, S.E., Anson, B.J., Mesecar, A.D.(2016) Sci Rep 6: 25961-25961
- PubMed: 27173881 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep25961
- Primary Citation Related Structures:
5HYO - PubMed Abstract:
Porcine epidemic diarrhea virus (PEDV) is a coronavirus that infects pigs and can have mortality rates approaching 100% in piglets, causing serious economic impact. The 3C-like protease (3CL(pro)) is essential for the coronaviral life cycle and is an appealing target for the development of therapeutics. We report the expression, purification, crystallization and 2.10 Å X-ray structure of 3CL(pro) from PEDV. Analysis of the PEDV 3CL(pro) structure and comparison to other coronaviral 3CL(pro)'s from the same alpha-coronavirus phylogeny shows that the overall structures and active site architectures across 3CL(pro)'s are conserved, with the exception of a loop that comprises the protease S2 pocket. We found a known inhibitor of severe acute respiratory syndrome coronavirus (SARS-CoV) 3CL(pro), (R)-16, to have inhibitor activity against PEDV 3CL(pro), despite that SARS-3CL(pro) and PEDV 3CL(pro) share only 45.4% sequence identity. Structural comparison reveals that the majority of residues involved in (R)-16 binding to SARS-3CL(pro) are conserved in PEDV-3CL(pro); however, the sequence variation and positional difference in the loop forming the S2 pocket may account for large observed difference in IC50 values. This work advances our understanding of the subtle, but important, differences in coronaviral 3CL(pro) architecture and contributes to the broader structural knowledge of coronaviral 3CL(pro)'s.
- Department of Chemistry, Purdue University, West Lafayette, Indiana, USA.
Organizational Affiliation: