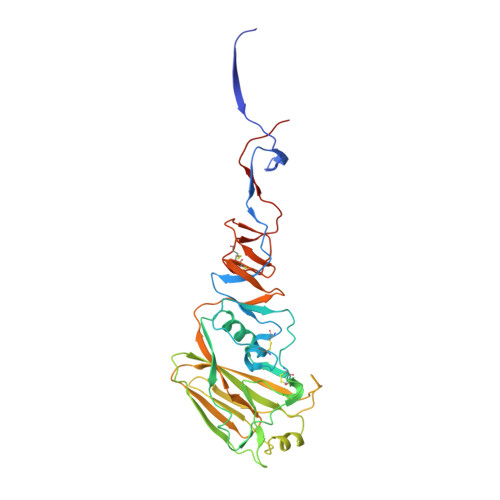
Molecular Characterizations of Surface Proteins Hemagglutinin and Neuraminidase from Recent H5Nx Avian Influenza Viruses.
Yang, H., Carney, P.J., Mishin, V.P., Guo, Z., Chang, J.C., Wentworth, D.E., Gubareva, L.V., Stevens, J.(2016) J Virol 90: 5770-5784
- PubMed: 27053557 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00180-16
- Primary Citation Related Structures:
5HU8, 5HUF, 5HUG, 5HUK, 5HUM, 5HUN - PubMed Abstract:
During 2014, a subclade 2.3.4.4 highly pathogenic avian influenza (HPAI) A(H5N8) virus caused poultry outbreaks around the world. In late 2014/early 2015, the virus was detected in wild birds in Canada and the United States, and these viruses also gave rise to reassortant progeny, composed of viral RNA segments (vRNAs) from both Eurasian and North American lineages. In particular, viruses were found with N1, N2, and N8 neuraminidase vRNAs, and these are collectively referred to as H5Nx viruses. In the United States, more than 48 million domestic birds have been affected. Here we present a detailed structural and biochemical analysis of the surface antigens of H5N1, H5N2, and H5N8 viruses in addition to those of a recent human H5N6 virus. Our results with recombinant hemagglutinin reveal that these viruses have a strict avian receptor binding preference, while recombinantly expressed neuraminidases are sensitive to FDA-approved and investigational antivirals. Although H5Nx viruses currently pose a low risk to humans, it is important to maintain surveillance of these circulating viruses and to continually assess future changes that may increase their pandemic potential. The H5Nx viruses emerging in North America, Europe, and Asia pose a great public health concern. Here we report a molecular and structural study of the major surface proteins of several H5Nx influenza viruses. Our results improve the understanding of these new viruses and provide important information on their receptor preferences and susceptibilities to antivirals, which are central to pandemic risk assessment.
- Influenza Division, National Center for Immunization and Respiratory Diseases, Centers for Disease Control and Prevention, Atlanta, Georgia, USA.
Organizational Affiliation: