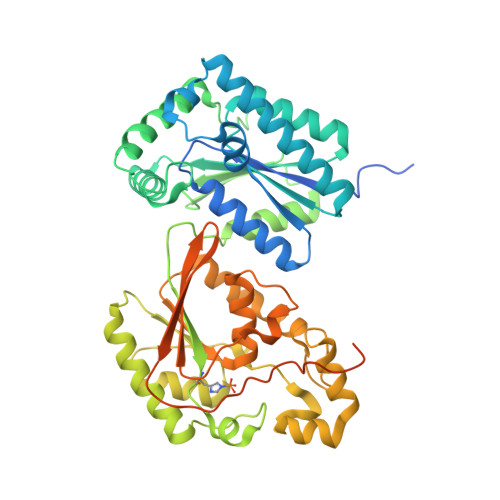
Crystal structure of heart 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase (PFKFB2) and the inhibitory influence of citrate on substrate binding.
Crochet, R.B., Kim, J.D., Lee, H., Yim, Y.S., Kim, S.G., Neau, D., Lee, Y.H.(2017) Proteins 85: 117-124
- PubMed: 27802586 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.25204
- Primary Citation Related Structures:
5HR5, 5HTK - PubMed Abstract:
The heart-specific isoform of 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase (PFKFB2) is an important regulator of glycolytic flux in cardiac cells. Here, we present the crystal structures of two PFKFB2 orthologues, human and bovine, at resolutions of 2.0 and 1.8 Å, respectively. Citrate, a TCA cycle intermediate and well-known inhibitor of PFKFB2, co-crystallized in the 2-kinase domains of both orthologues, occupying the fructose-6-phosphate binding-site and extending into the γ-phosphate binding pocket of ATP. This steric and electrostatic occlusion of the γ-phosphate site by citrate proved highly consequential to the binding of co-complexed ATP analogues. The bovine structure, which co-crystallized with ADP, closely resembled the overall structure of other PFKFB isoforms, with ADP mimicking the catalytic binding mode of ATP. The human structure, on the other hand, co-complexed with AMPPNP, which, unlike ADP, contains a γ-phosphate. The presence of this γ-phosphate made adoption of the catalytic ATP binding mode impossible for AMPPNP, forcing the analogue to bind atypically with concomitant conformational changes to the ATP binding-pocket. Inhibition kinetics were used to validate the structural observations, confirming citrate's inhibition mechanism as competitive for F6P and noncompetitive for ATP. Together, these structural and kinetic data establish a molecular basis for citrate's negative feed-back loop of the glycolytic pathway via PFKFB2. Proteins 2016; 85:117-124. © 2016 Wiley Periodicals, Inc.
- Departments of Biological Sciences, Louisiana State University, Baton Rouge, Louisiana, 70803.
Organizational Affiliation: