The crystal structure of Se-AsfvPolX(L52/163M mutant) in complex with 1nt-gap DNA1
Chen, Y.Q., Zhang, J., Gan, J.H.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA polymerase beta-like protein | 178 | African swine fever virus | Mutation(s): 2 Gene Names: O174L EC: 2.7.7.7 | 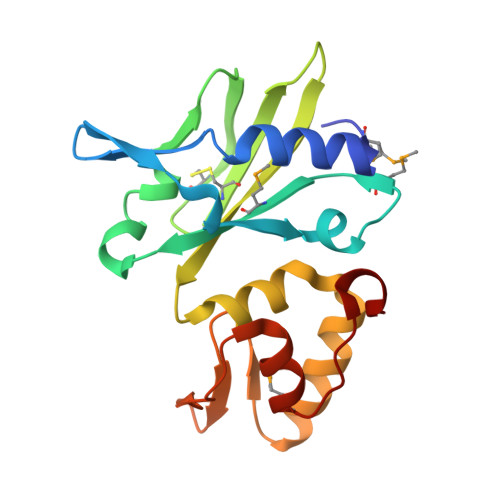 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P42494 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*GP*GP*AP*CP*AP*AP*CP*GP*GP*GP*AP*CP*AP*AP*C)-3') | 15 | synthetic construct | 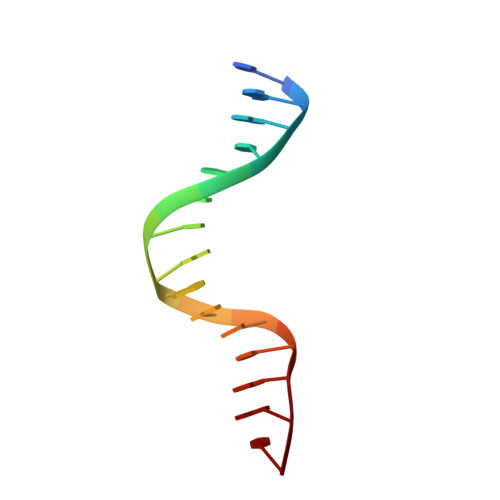 | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A, B | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 74.323 | α = 90 |
| b = 82.24 | β = 90 |
| c = 101.416 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Fundation of China | China | 31370728 |