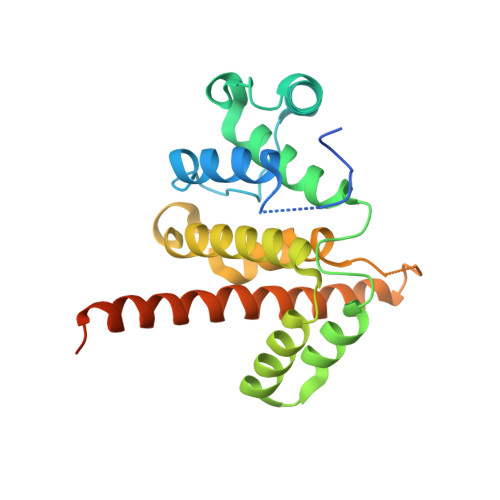
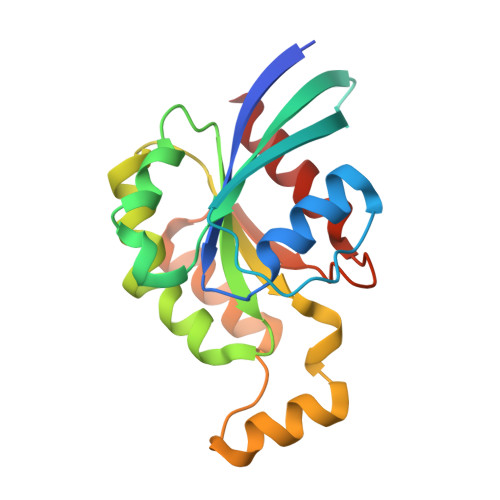
Noncanonical Myo9b-RhoGAP Accelerates RhoA GTP Hydrolysis by a Dual-Arginine-Finger Mechanism
Yi, F., Kong, R., Ren, J., Zhu, L., Lou, J., Wu, J.Y., Feng, W.(2016) J Mol Biology 428: 3043-3057
- PubMed: 27363609 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2016.06.014
- Primary Citation Related Structures:
5HPY - PubMed Abstract:
The GTP hydrolysis activities of Rho GTPases are stimulated by GTPase-activating proteins (GAPs), which contain a RhoGAP domain equipped with a characteristic arginine finger and an auxiliary asparagine for catalysis. However, the auxiliary asparagine is missing in the RhoGAP domain of Myo9b (Myo9b-RhoGAP), a unique motorized RhoGAP that specifically targets RhoA for controlling cell motility. Here, we determined the structure of Myo9b-RhoGAP in complex with GDP-bound RhoA and magnesium fluoride. Unexpectedly, Myo9b-RhoGAP contains two arginine fingers at its catalytic site. The first arginine finger resembles the one within the canonical RhoGAP domains and inserts into the nucleotide-binding pocket of RhoA, whereas the second arginine finger anchors the Switch I loop of RhoA and interacts with the nucleotide, stabilizing the transition state of GTP hydrolysis and compensating for the lack of the asparagine. Mutating either of the two arginine fingers impaired the catalytic activity of Myo9b-RhoGAP and affected the Myo9b-mediated cell migration. Our data indicate that Myo9b-RhoGAP accelerates RhoA GTP hydrolysis by a previously unknown dual-arginine-finger mechanism, which may be shared by other noncanonical RhoGAP domains lacking the auxiliary asparagine.
- National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Beijing 100101, China; University of Chinese Academy of Sciences, Beijing 100049, China.
Organizational Affiliation: