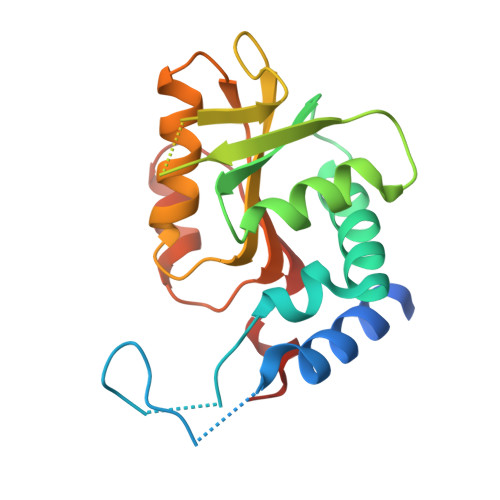
Structural investigations on orotate phosphoribosyltransferase from Mycobacterium tuberculosis, a key enzyme of the de novo pyrimidine biosynthesis.
Donini, S., Ferraris, D.M., Miggiano, R., Massarotti, A., Rizzi, M.(2017) Sci Rep 7: 1180-1180
- PubMed: 28446777 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-01057-z
- Primary Citation Related Structures:
5HKF, 5HKI, 5HKL - PubMed Abstract:
The Mycobacterium tuberculosis orotate phosphoribosyltransferase (MtOPRT) catalyses the conversion of α-D-5-phosphoribosyl-1-pyrophosphate (PRPP) and orotate (OA) in pyrophosphate and orotidine 5'-monophosphate (OMP), in presence of Mg 2+ . This enzyme is the only responsible for the synthesis of orotidine 5'-monophosphate, a key precursor in the de novo pyrimidine biosynthesis pathway, making MtOPRT an attractive drug target for the development of antitubercular agents. We report the crystal structures of MtOPRT in complex with PRPP (2.25 Å resolution), inorganic phosphate (1.90 Å resolution) and the exogenous compound Fe(III) dicitrate (2.40 Å resolution). The overall structure of the mycobacterial enzyme is highly similar to those described for other OPRTases, with the "flexible loop" assuming a well define conformation and making specific contacts with the Fe(III)-dicitrate complex. The structures here reported add to the knowledge of a potential drug target for tuberculosis, and will provide a useful tool for the structure-based drug design of potent enzyme inhibitors.
- Department of Pharmaceutical Sciences, Università del Piemonte Orientale "A. Avogadro", Largo Donegani 2, 28100, Novara, Italy.
Organizational Affiliation: