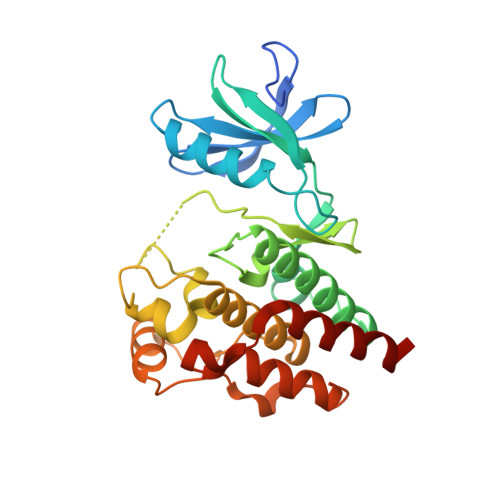
Activation Mechanism of Oncogenic Deletion Mutations in BRAF, EGFR, and HER2.
Foster, S.A., Whalen, D.M., Ozen, A., Wongchenko, M.J., Yin, J., Yen, I., Schaefer, G., Mayfield, J.D., Chmielecki, J., Stephens, P.J., Albacker, L.A., Yan, Y., Song, K., Hatzivassiliou, G., Eigenbrot, C., Yu, C., Shaw, A.S., Manning, G., Skelton, N.J., Hymowitz, S.G., Malek, S.(2016) Cancer Cell 29: 477-493
- PubMed: 26996308 Search on PubMed
- DOI: https://doi.org/10.1016/j.ccell.2016.02.010
- Primary Citation Related Structures:
5HI2, 5HIB, 5HIC, 5HID, 5HIE - PubMed Abstract:
Activating mutations in protein kinases drive many cancers. While how recurring point mutations affect kinase activity has been described, the effect of in-frame deletions is not well understood. We show that oncogenic deletions within the β3-αC loop of HER2 and BRAF are analogous to the recurrent EGFR exon 19 deletions. We identify pancreatic carcinomas with BRAF deletions mutually exclusive with KRAS mutations. Crystal structures of BRAF deletions reveal the truncated loop restrains αC in an active "in" conformation, imparting resistance to inhibitors like vemurafenib that bind the αC "out" conformation. Characterization of loop length explains the prevalence of five amino acid deletions in BRAF, EGFR, and HER2 and highlights the importance of this region for kinase activity and inhibitor efficacy.
- Department of Discovery Oncology, Genentech Inc., 1 DNA Way, South San Francisco, CA 94080, USA.
Organizational Affiliation: