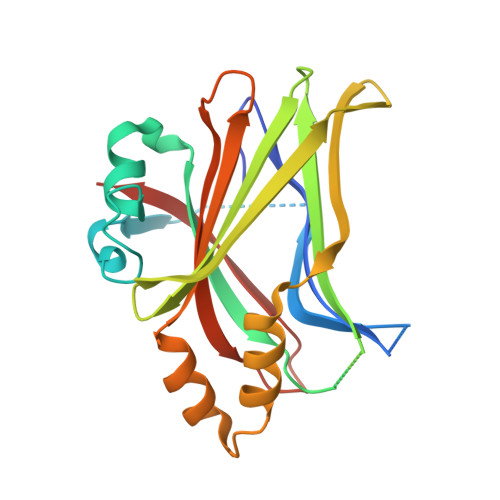
Autopalmitoylation of TEAD proteins regulates transcriptional output of the Hippo pathway.
Chan, P., Han, X., Zheng, B., DeRan, M., Yu, J., Jarugumilli, G.K., Deng, H., Pan, D., Luo, X., Wu, X.(2016) Nat Chem Biol 12: 282-289
- PubMed: 26900866 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.2036
- Primary Citation Related Structures:
5HGU - PubMed Abstract:
TEA domain (TEAD) transcription factors bind to the coactivators YAP and TAZ and regulate the transcriptional output of the Hippo pathway, playing critical roles in organ size control and tumorigenesis. Protein S-palmitoylation attaches a fatty acid, palmitate, to cysteine residues and regulates protein trafficking, membrane localization and signaling activities. Using activity-based chemical probes, we discovered that human TEADs possess intrinsic palmitoylating enzyme-like activities and undergo autopalmitoylation at evolutionarily conserved cysteine residues under physiological conditions. We determined the crystal structures of lipid-bound TEADs and found that the lipid chain of palmitate inserts into a conserved deep hydrophobic pocket. Strikingly, palmitoylation did not alter TEAD's localization, but it was required for TEAD's binding to YAP and TAZ and was dispensable for its binding to the Vgll4 tumor suppressor. Moreover, palmitoylation-deficient TEAD mutants impaired TAZ-mediated muscle differentiation in vitro and tissue overgrowth mediated by the Drosophila YAP homolog Yorkie in vivo. Our study directly links autopalmitoylation to the transcriptional regulation of the Hippo pathway.
- Cutaneous Biology Research Center, Massachusetts General Hospital, Harvard Medical School, Charlestown, Massachusetts, USA.
Organizational Affiliation: