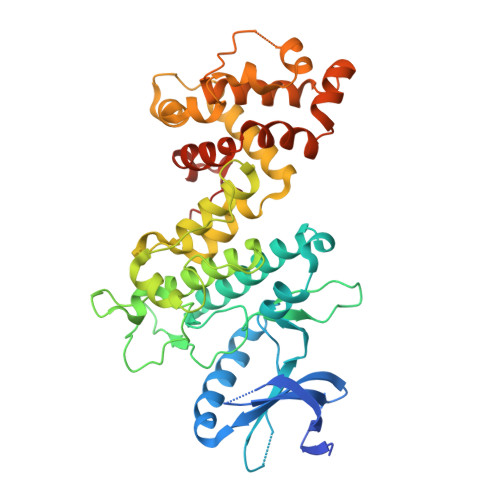
Structural and Functional Analysis of the Allosteric Inhibition of IRE1 alpha with ATP-Competitive Ligands.
Feldman, H.C., Tong, M., Wang, L., Meza-Acevedo, R., Gobillot, T.A., Lebedev, I., Gliedt, M.J., Hari, S.B., Mitra, A.K., Backes, B.J., Papa, F.R., Seeliger, M.A., Maly, D.J.(2016) ACS Chem Biol 11: 2195-2205
- PubMed: 27227314 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.5b00940
- Primary Citation Related Structures:
5HGI - PubMed Abstract:
The accumulation of unfolded proteins under endoplasmic reticulum (ER) stress leads to the activation of the multidomain protein sensor IRE1α as part of the unfolded protein response (UPR). Clustering of IRE1α lumenal domains in the presence of unfolded proteins promotes kinase trans-autophosphorylation in the cytosol and subsequent RNase domain activation. Interestingly, there is an allosteric relationship between the kinase and RNase domains of IRE1α, which allows ATP-competitive inhibitors to modulate the activity of the RNase domain. Here, we use kinase inhibitors to study how ATP-binding site conformation affects the activity of the RNase domain of IRE1α. We find that diverse ATP-competitive inhibitors of IRE1α promote dimerization and activation of RNase activity despite blocking kinase autophosphorylation. In contrast, a subset of ATP-competitive ligands, which we call KIRAs, allosterically inactivate the RNase domain through the kinase domain by stabilizing monomeric IRE1α. Further insight into how ATP-competitive inhibitors are able to divergently modulate the RNase domain through the kinase domain was gained by obtaining the first structure of apo human IRE1α in the RNase active back-to-back dimer conformation. Comparison of this structure with other existing structures of IRE1α and integration of our extensive structure activity relationship (SAR) data has led us to formulate a model to rationalize how ATP-binding site ligands are able to control the IRE1α oligomeric state and subsequent RNase domain activity.
- Department of Chemistry, University of Washington , Seattle, Washington, United States.
Organizational Affiliation: