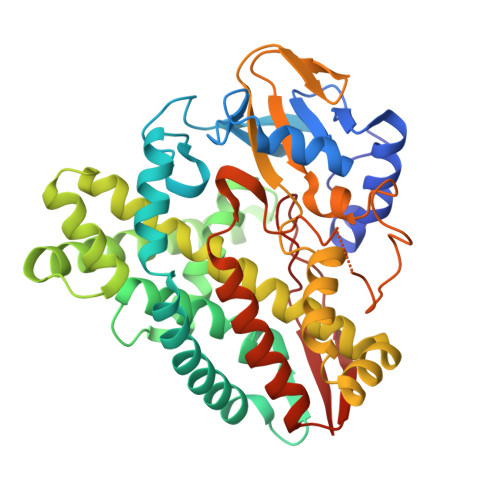
Structural characterization of CYP144A1 - a cytochrome P450 enzyme expressed from alternative transcripts in Mycobacterium tuberculosis.
Chenge, J., Kavanagh, M.E., Driscoll, M.D., McLean, K.J., Young, D.B., Cortes, T., Matak-Vinkovic, D., Levy, C.W., Rigby, S.E., Leys, D., Abell, C., Munro, A.W.(2016) Sci Rep 6: 26628-26628
- PubMed: 27225995 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep26628
- Primary Citation Related Structures:
5HDI - PubMed Abstract:
Mycobacterium tuberculosis (Mtb) causes the disease tuberculosis (TB). The virulent Mtb H37Rv strain encodes 20 cytochrome P450 (CYP) enzymes, many of which are implicated in Mtb survival and pathogenicity in the human host. Bioinformatics analysis revealed that CYP144A1 is retained exclusively within the Mycobacterium genus, particularly in species causing human and animal disease. Transcriptomic annotation revealed two possible CYP144A1 start codons, leading to expression of (i) a "full-length" 434 amino acid version (CYP144A1-FLV) and (ii) a "truncated" 404 amino acid version (CYP144A1-TRV). Computational analysis predicted that the extended N-terminal region of CYP144A1-FLV is largely unstructured. CYP144A1 FLV and TRV forms were purified in heme-bound states. Mass spectrometry confirmed production of intact, His6-tagged forms of CYP144A1-FLV and -TRV, with EPR demonstrating cysteine thiolate coordination of heme iron in both cases. Hydrodynamic analysis indicated that both CYP144A1 forms are monomeric. CYP144A1-TRV was crystallized and the first structure of a CYP144 family P450 protein determined. CYP144A1-TRV has an open structure primed for substrate binding, with a large active site cavity. Our data provide the first evidence that Mtb produces two different forms of CYP144A1 from alternative transcripts, with CYP144A1-TRV generated from a leaderless transcript lacking a 5'-untranslated region and Shine-Dalgarno ribosome binding site.
- Manchester Institute of Biotechnology, Centre for Synthetic Biology of Fine and Specialty Chemicals (SYNBIOCHEM), Faculty of Life Sciences, The University of Manchester, Manchester M1 7DN, United Kingdom.
Organizational Affiliation: