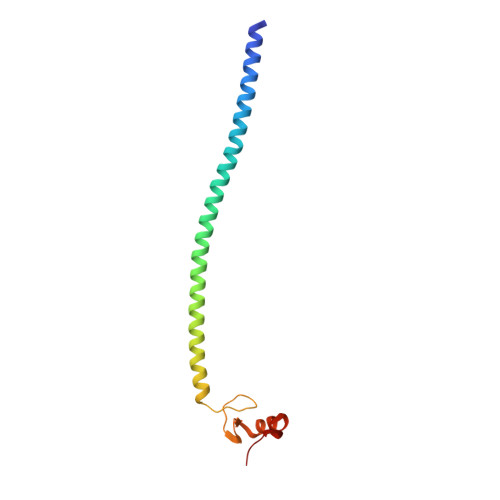
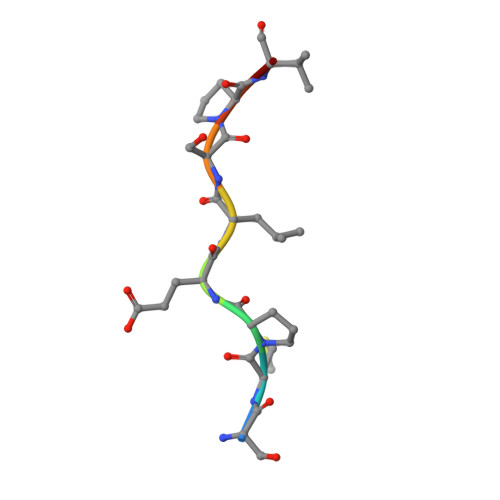
BS69/ZMYND11 C-Terminal Domains Bind and Inhibit EBNA2.
Harter, M.R., Liu, C.D., Shen, C.L., Gonzalez-Hurtado, E., Zhang, Z.M., Xu, M., Martinez, E., Peng, C.W., Song, J.(2016) PLoS Pathog 12: e1005414-e1005414
- PubMed: 26845565 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1005414
- Primary Citation Related Structures:
5HDA - PubMed Abstract:
Epstein-Barr virus (EBV) nuclear antigen 2 (EBNA2) plays an important role in driving immortalization of EBV-infected B cells through regulating the expression of many viral and cellular genes. We report a structural study of the tumor suppressor BS69/ZMYND11 C-terminal region, comprised of tandem coiled-coil-MYND domains (BS69CC-MYND), in complex with an EBNA2 peptide containing a PXLXP motif. The coiled-coil domain of BS69 self-associates to bring two separate MYND domains in close proximity, thereby enhancing the BS69 MYND-EBNA2 interaction. ITC analysis of BS69CC-MYND with a C-terminal fragment of EBNA2 further suggests that the BS69CC-MYND homodimer synergistically binds to the two EBNA2 PXLXP motifs that are respectively located in the conserved regions CR7 and CR8. Furthermore, we showed that EBNA2 interacts with BS69 and down-regulates its expression at both mRNA and protein levels in EBV-infected B cells. Ectopic BS69CC-MYND is recruited to viral target promoters through interactions with EBNA2, inhibits EBNA2-mediated transcription activation, and impairs proliferation of lymphoblastoid cell lines (LCLs). Substitution of critical residues in the MYND domain impairs the BS69-EBNA2 interaction and abolishes the BS69 inhibition of the EBNA2-mediated transactivation and LCL proliferation. This study identifies the BS69 C-terminal domains as an inhibitor of EBNA2, which may have important implications in development of novel therapeutic strategies against EBV infection.
- Department of Biochemistry, University of California, Riverside, Riverside, California, United States of America.
Organizational Affiliation: