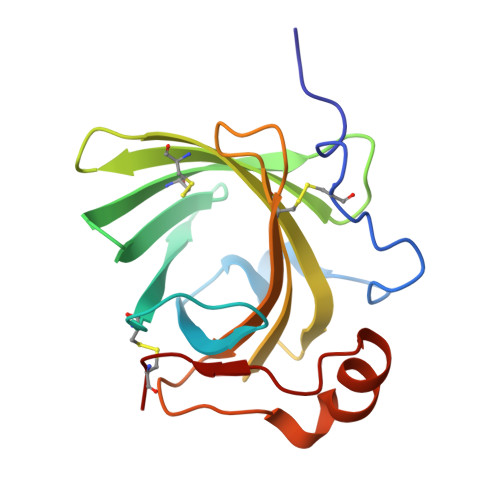
Structure and Ligand-Binding Mechanism of a Cysteinyl Leukotriene-Binding Protein from a Blood-Feeding Disease Vector.
Jablonka, W., Pham, V., Nardone, G., Gittis, A., Silva-Cardoso, L., Atella, G.C., Ribeiro, J.M., Andersen, J.F.(2016) ACS Chem Biol 11: 1934-1944
- PubMed: 27124118 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.6b00032
- Primary Citation Related Structures:
5H9K, 5H9L, 5H9N, 5HA0, 5HAE - PubMed Abstract:
Blood-feeding disease vectors mitigate the negative effects of hemostasis and inflammation through the binding of small-molecule agonists of these processes by salivary proteins. In this study, a lipocalin protein family member (LTBP1) from the saliva of Rhodnius prolixus, a vector of the pathogen Trypanosoma cruzi, is shown to sequester cysteinyl leukotrienes during feeding to inhibit immediate inflammatory responses. Calorimetric binding experiments showed that LTBP1 binds leukotrienes C4 (LTC4), D4 (LTD4), and E4 (LTE4) but not biogenic amines, adenosine diphosphate, or other eicosanoid compounds. Crystal structures of ligand-free LTBP1 and its complexes with LTC4 and LTD4 reveal a conformational change during binding that brings Tyr114 into close contact with the ligand. LTC4 is cleaved in the complex, leaving free glutathione and a C20 fatty acid. Chromatographic analysis of bound ligands showed only intact LTC4, suggesting that cleavage could be radiation-mediated.
- Laboratory of Malaria and Vector Research, NIAID, National Institutes of Health , Rockville, Maryland 20852, United States.
Organizational Affiliation: