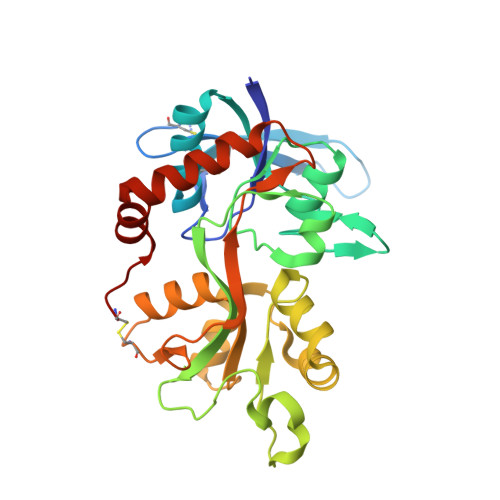
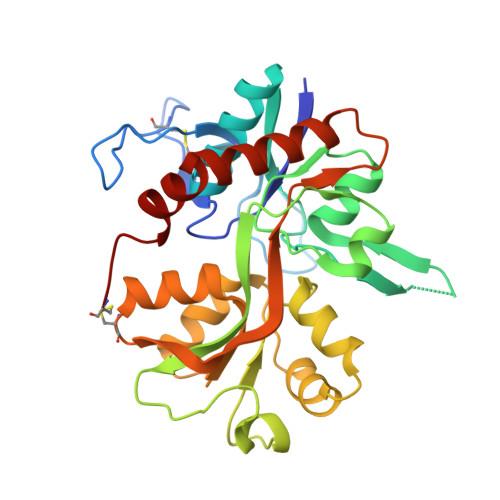
Positive Allosteric Modulators of GluN2A-Containing NMDARs with Distinct Modes of Action and Impacts on Circuit Function.
Hackos, D.H., Lupardus, P.J., Grand, T., Chen, Y., Wang, T.M., Reynen, P., Gustafson, A., Wallweber, H.J., Volgraf, M., Sellers, B.D., Schwarz, J.B., Paoletti, P., Sheng, M., Zhou, Q., Hanson, J.E.(2016) Neuron 89: 983-999
- PubMed: 26875626 Search on PubMed
- DOI: https://doi.org/10.1016/j.neuron.2016.01.016
- Primary Citation Related Structures:
5H8F, 5H8H, 5H8N, 5H8Q, 5H8S, 5KCJ - PubMed Abstract:
To enhance physiological function of NMDA receptors (NMDARs), we identified positive allosteric modulators (PAMs) of NMDARs with selectivity for GluN2A subunit-containing receptors. X-ray crystallography revealed a binding site at the GluN1-GluN2A dimer interface of the extracellular ligand-binding domains (LBDs). Despite the similarity between the LBDs of NMDARs and AMPA receptors (AMPARs), GluN2A PAMs with good selectivity against AMPARs were identified. Potentiation was observed with recombinant triheteromeric GluN1/GluN2A/GluN2B NMDARs and with synaptically activated NMDARs in brain slices from wild-type (WT), but not GluN2A knockout (KO), mice. Individual GluN2A PAMs exhibited variable degrees of glutamate (Glu) dependence, impact on NMDAR Glu EC50, and slowing of channel deactivation. These distinct PAMs also exhibited differential impacts during synaptic plasticity induction. The identification of a new NMDAR modulatory site and characterization of GluN2A-selective PAMs provide powerful molecular tools to dissect NMDAR function and demonstrate the feasibility of a therapeutically desirable type of NMDAR enhancement.
- Department of Neuroscience, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA.
Organizational Affiliation: