Mechanistic insights into the alternative translation termination by ArfA and RF2
Ma, C., Kurita, D., Li, N., Chen, Y., Himeno, H., Gao, N.(2017) Nature 541: 550-553
- PubMed: 27906160 Search on PubMed
- DOI: https://doi.org/10.1038/nature20822
- Primary Citation Related Structures:
5H5U - PubMed Abstract:
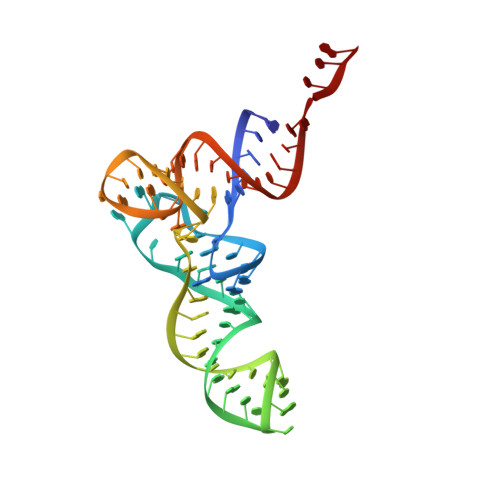
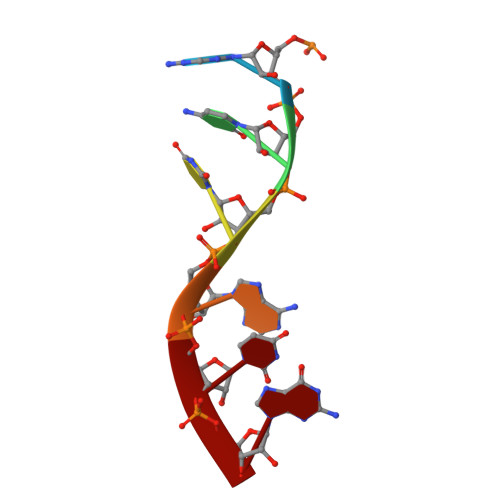
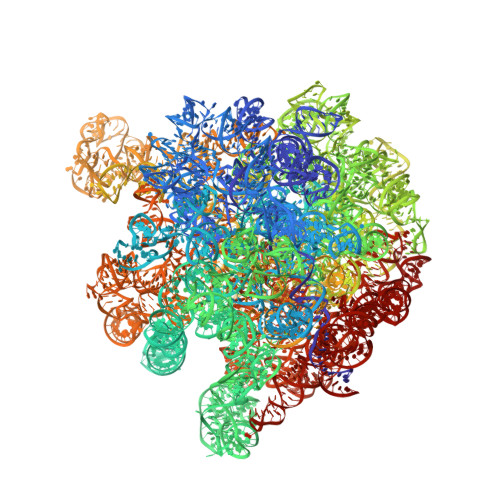
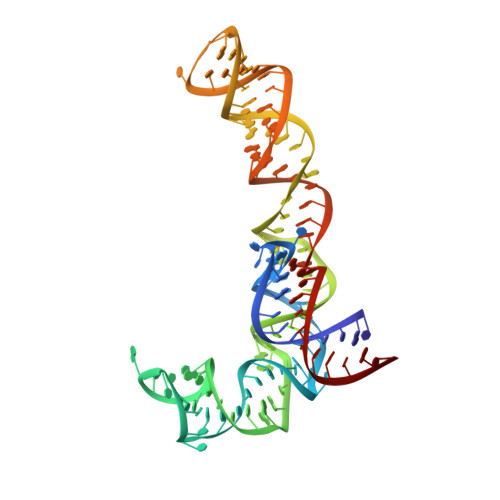
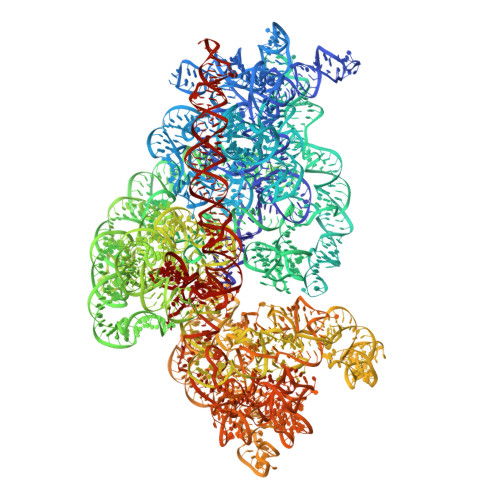
During cellular translation of messenger RNAs by ribosomes, the translation apparatus sometimes pauses or stalls at the elongation and termination steps. With the exception of programmed stalling, which is usually used by cells for regulatory purposes, ribosomes stalled on mRNAs need to be terminated and recycled to maintain adequate translation capacity. Much ribosome stalling originates in aberrant mRNAs that lack a stop codon. Transcriptional errors, misprocessing of primary transcripts, and undesired mRNA cleavage all contribute to the formation of non-stop mRNAs. Ribosomes stalled at the 3' end of non-stop mRNAs do not undergo normal termination owing to the lack of specific stop-codon recognition by canonical peptide release factors at the A-site decoding centre. In bacteria, the transfer-messenger RNA (tmRNA)-SmpB-mediated trans-translation rescue system reroutes stalled ribosomes to the normal elongation cycle and translation termination. Two additional rescue systems, ArfA-RF2 (refs 13, 14, 15, 16) and ArfB (formerly known as YaeJ), are also present in many bacterial species, but their mechanisms are not fully understood. Here, using cryo-electron microscopy, we characterize the structure of the Escherichia coli 70S ribosome bound with ArfA, the release factor RF2, a short non-stop mRNA and a cognate P-site tRNA. The C-terminal loop of ArfA occupies the mRNA entry channel on the 30S subunit, whereas its N terminus is sandwiched between the decoding centre and the switch loop of RF2, leading to marked conformational changes in both the decoding centre and RF2. Despite the distinct conformation of RF2, its conserved catalytic GGQ motif is precisely positioned next to the CCA-end of the P-site tRNA. These data illustrate a stop-codon surrogate mechanism for ArfA in facilitating the termination of non-stop ribosomal complexes by RF2.
- Ministry of Education Key Laboratory of Protein Sciences, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: