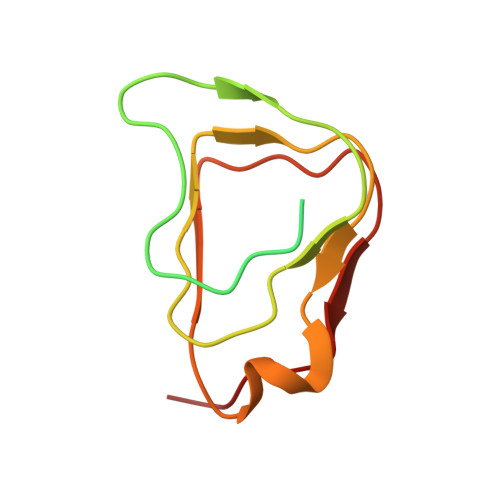
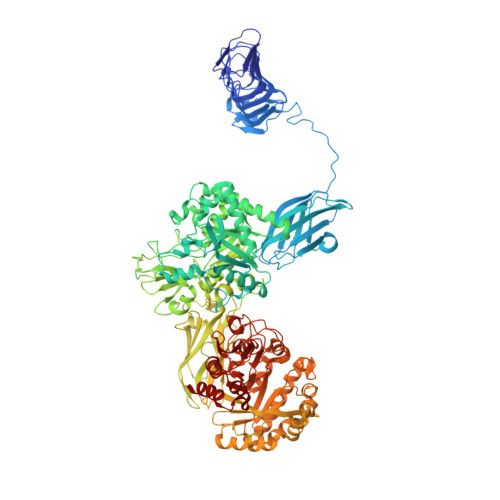
Crystal Structure of Chitinase ChiW from Paenibacillus sp. str. FPU-7 Reveals a Novel Type of Bacterial Cell-Surface-Expressed Multi-Modular Enzyme Machinery
Itoh, T., Hibi, T., Suzuki, F., Sugimoto, I., Fujiwara, A., Inaka, K., Tanaka, H., Ohta, K., Fujii, Y., Taketo, A., Kimoto, H.(2016) PLoS One 11: e0167310-e0167310
- PubMed: 27907169 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0167310
- Primary Citation Related Structures:
5GZT, 5GZU, 5GZV - PubMed Abstract:
The Gram-positive bacterium Paenibacillus sp. str. FPU-7 effectively hydrolyzes chitin by using a number of chitinases. A unique chitinase with two catalytic domains, ChiW, is expressed on the cell surface of this bacterium and has high activity towards various chitins, even crystalline chitin. Here, the crystal structure of ChiW at 2.1 Å resolution is presented and describes how the enzyme degrades chitin on the bacterial cell surface. The crystal structure revealed a unique multi-modular architecture composed of six domains to function efficiently on the cell surface: a right-handed β-helix domain (carbohydrate-binding module family 54, CBM-54), a Gly-Ser-rich loop, 1st immunoglobulin-like (Ig-like) fold domain, 1st β/α-barrel catalytic domain (glycoside hydrolase family 18, GH-18), 2nd Ig-like fold domain and 2nd β/α-barrel catalytic domain (GH-18). The structure of the CBM-54, flexibly linked to the catalytic region of ChiW, is described here for the first time. It is similar to those of carbohydrate lyases but displayed no detectable carbohydrate degradation activities. The CBM-54 of ChiW bound to cell wall polysaccharides, such as chin, chitosan, β-1,3-glucan, xylan and cellulose. The structural and biochemical data obtained here also indicated that the enzyme has deep and short active site clefts with endo-acting character. The affinity of CBM-54 towards cell wall polysaccharides and the degradation pattern of the catalytic domains may help to efficiently decompose the cell wall chitin through the contact surface. Furthermore, we clarify that other Gram-positive bacteria possess similar cell-surface-expressed multi-modular enzymes for cell wall polysaccharide degradation.
- Department of Bioscience, Fukui Prefectural University, Yoshida-gun, Fukui, Japan.
Organizational Affiliation: