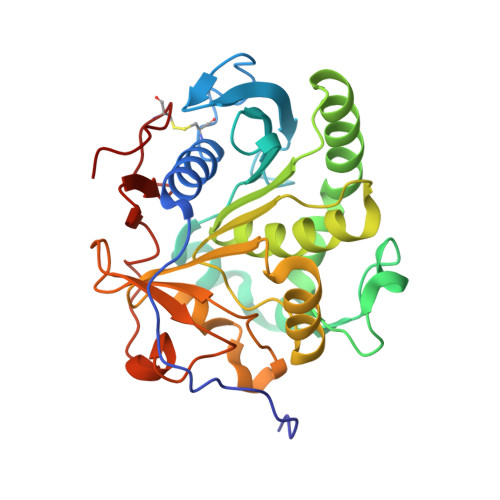
Malassezia globosa MgMDL2 lipase: Crystal structure and rational modification of substrate specificity.
Lan, D., Xu, H., Xu, J., Dubin, G., Liu, J., Iqbal Khan, F., Wang, Y.(2017) Biochem Biophys Res Commun 488: 259-265
- PubMed: 28433636 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.04.103
- Primary Citation Related Structures:
5GW8 - PubMed Abstract:
Lipases play an important role in physiological metabolism and diseases, and also have multiple industrial applications. Rational modification of lipase specificity may increase the commercial utility of this group of enzymes, but is hindered by insufficient mechanistic understanding. Here, we report the 2.0 Å resolution crystal structure of a mono- and di-acylglycerols lipase from Malassezia globosa (MgMDL2). Interestingly, residues Phe278 and Glu282 were found to involve in substrate recognition because mutation on each residue led to convert MgMDL2 to a triacylglycerol (TAG) lipase. The Phe278Ala and Glu282Ala mutants also acquired ability to synthesize TAGs by esterification of glycerol and fatty acids. By in silicon analysis, steric hindrance of these residues seemed to be key factors for the altered substrate specificity. Our work may shed light on understanding the unique substrate selectivity mechanism of mono- and di-acylglycerols lipases, and provide a new insight for engineering biocatalysts with desired catalytic behaviors for biotechnological application.
- School of Food Science and Engineering, South China University of Technology, Guangzhou, PR China.
Organizational Affiliation: