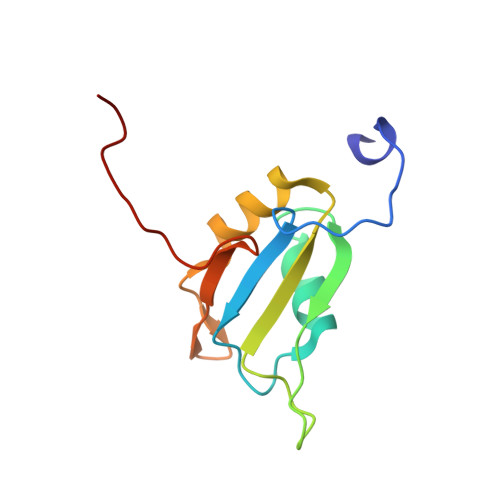
Solution structure of the first RNA recognition motif domain of human spliceosomal protein SF3b49 and its mode of interaction with a SF3b145 fragment.
Kuwasako, K., Nameki, N., Tsuda, K., Takahashi, M., Sato, A., Tochio, N., Inoue, M., Terada, T., Kigawa, T., Kobayashi, N., Shirouzu, M., Ito, T., Sakamoto, T., Wakamatsu, K., Guntert, P., Takahashi, S., Yokoyama, S., Muto, Y.(2017) Protein Sci 26: 280-291
- PubMed: 27862552 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3080
- Primary Citation Related Structures:
5GVQ - PubMed Abstract:
The spliceosomal protein SF3b49, a component of the splicing factor 3b (SF3b) protein complex in the U2 small nuclear ribonucleoprotein, contains two RNA recognition motif (RRM) domains. In yeast, the first RRM domain (RRM1) of Hsh49 protein (yeast orthologue of human SF3b49) reportedly interacts with another component, Cus1 protein (orthologue of human SF3b145). Here, we solved the solution structure of the RRM1 of human SF3b49 and examined its mode of interaction with a fragment of human SF3b145 using NMR methods. Chemical shift mapping showed that the SF3b145 fragment spanning residues 598-631 interacts with SF3b49 RRM1, which adopts a canonical RRM fold with a topology of β1-α1-β2-β3-α2-β4. Furthermore, a docking model based on NOESY measurements suggests that residues 607-616 of the SF3b145 fragment adopt a helical structure that binds to RRM1 predominantly via α1, consequently exhibiting a helix-helix interaction in almost antiparallel. This mode of interaction was confirmed by a mutational analysis using GST pull-down assays. Comparison with structures of all RRM domains when complexed with a peptide found that this helix-helix interaction is unique to SF3b49 RRM1. Additionally, all amino acid residues involved in the interaction are well conserved among eukaryotes, suggesting evolutionary conservation of this interaction mode between SF3b49 RRM1 and SF3b145.
- Faculty of Pharmacy and Research Institute of Pharmaceutical Sciences, Musashino University, 1-1-20 Shinmachi, Nishitokyo, Tokyo, 202-8585, Japan.
Organizational Affiliation: