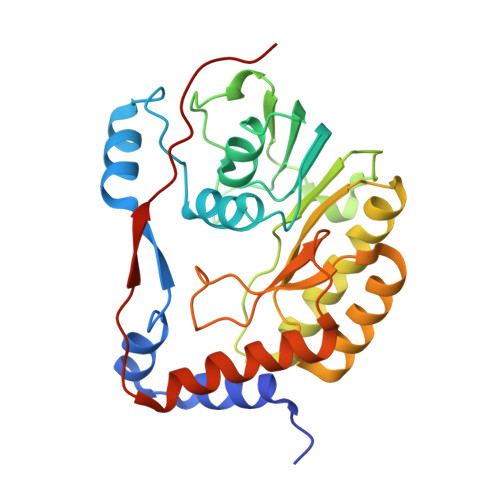
Structure of the NS5 methyltransferase from Zika virus and implications in inhibitor design
Zhang, C., Feng, T., Cheng, J., Li, Y., Yin, X., Zeng, W., Jin, X., Li, Y., Guo, F., Jin, T.(2017) Biochem Biophys Res Commun 492: 624-630
- PubMed: 27866982 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2016.11.098
- Primary Citation Related Structures:
5GOZ, 5GP1 - PubMed Abstract:
Recent outbreak of flavivirus Zika virus (ZIKV) in America has urged the basic as well as translational studies of this important human pathogen. The nonstructural protein 5 (NS5) of the flavivirus has an N-terminal methyltransferase (MTase) domain that plays critical roles in viral RNA genome capping. The null mutant of NS5 MTase is lethal for virus. Therefore, NS5 is a potential drug target for the treatment of Zika virus infection. In this study, we determined crystal structures of the ZIKV MTase in complex with GTP and RNA cap analogue 7me GpppA. Structural analyses revealed highly conserved GTP/cap-binding pocket and S-adenosylmethionine (SAM)-binding pocket. Two conformations of the second base of the cap were identified, which suggests the flexibility of RNA conformation. In addition, the ligand-binding pockets identified a continuous region of hotspots suitable for drug design. Docking calculation shows that the Dengue virus inhibitor compound 10 may bind to the ZIKV MTase.
- Laboratory of Structural Immunology, CAS Key Laboratory of Innate Immunity and Chronic Disease, CAS Center for Excellence in Molecular Cell Science, School of Life Sciences and Medical Center, University of Science and Technology of China, Hefei, Anhui, 230027, China.
Organizational Affiliation: