Discovery of (R)-1-(3-(4-Amino-3-(3-chloro-4-(pyridin-2-ylmethoxy)phenyl)-1H-pyrazolo[3,4-d]pyrimidin-1-yl)piperidin-1-yl)prop-2-en-1-one (CHMFL-EGFR-202) as a Novel Irreversible EGFR Mutant Kinase Inhibitor with a Distinct Binding Mode.
Wang, A., Li, X., Wu, H., Zou, F., Yan, X.E., Chen, C., Hu, C., Yu, K., Wang, W., Zhao, P., Wu, J., Qi, Z., Wang, W., Wang, B., Wang, L., Ren, T., Zhang, S., Yun, C.H., Liu, J., Liu, Q.(2017) J Med Chem 60: 2944-2962
- PubMed: 28282122 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01907
- Primary Citation Related Structures:
5GNK - PubMed Abstract:
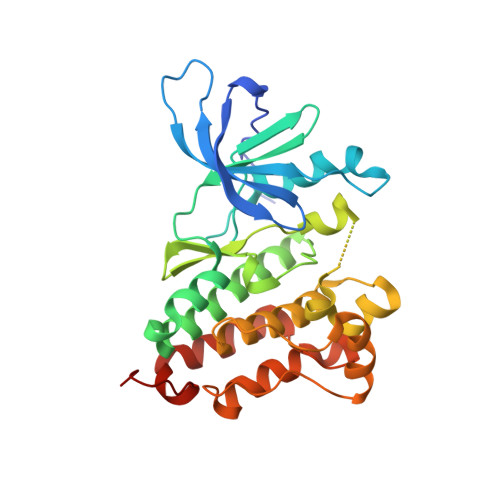
On the basis of Ibrutinib's core pharmacophore, which was moderately active to EGFR T790M mutant, we discovered novel epidermal growth factor receptor (EGFR) inhibitor compound 19 (CHMFL-EGFR-202), which potently inhibited EGFR primary mutants (L858R, del19) and drug-resistant mutant L858R/T790M. Compound 19 displayed a good selectivity profile among 468 kinases/mutants tested in the KINOMEscan assay (S score (1) = 0.02). In particular, it did not exhibit apparent activities against INSR and IGF1R kinases. The X-ray crystal structure revealed that this class of inhibitors formed a covalent bond with Cys797 in a distinct "DFG-in-C-helix-out" inactive EGFR conformation. Compound 19 displayed strong antiproliferative effects against EGFR mutant-driven nonsmall cell lung cancer (NSCLC) cell lines such as H1975, PC9, HCC827, and H3255 but not the wild-type EGFR expressing cells. In the H1975 and PC9 cell-inoculated xenograft mouse models, compound 19 exhibited dose-dependent tumor growth suppression efficacy without obvious toxicity. Compound 19 might be a potential drug candidate for EGFR mutant-driven NSCLC.
- High Magnetic Field Laboratory, Chinese Academy of Sciences , Mailbox 1110, 350 Shushanhu Road, Hefei, Anhui 230031, P. R. China.
Organizational Affiliation: