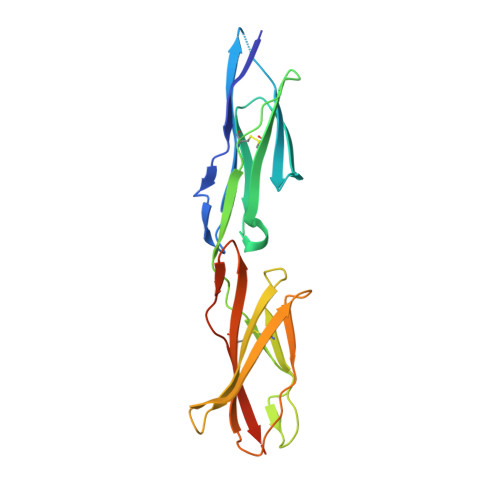
Structural Basis for Human PECAM-1-Mediated Trans-homophilic Cell Adhesion
Hu, M., Zhang, H., Liu, Q., Hao, Q.(2016) Sci Rep 6: 38655-38655
- PubMed: 27958302
- DOI: https://doi.org/10.1038/srep38655
- Primary Citation Related Structures:
5GNI - PubMed Abstract:
Cell adhesion involved in signal transduction, tissue integrity and pathogen infection is mainly mediated by cell adhesion molecules (CAM). One CAM member, platelet-endothelial-cell adhesion molecule-1 (PECAM-1), plays an important role in tight junction among endothelia cells, leukocyte trafficking, and immune response through its homophilic and heterophilic binding patterns. Both kinds of interactions, which lead to endogenous and exogenous signal transmission, are derived from extracellular immunoglobulin-like (IgL) domains and cytoplasmic immunoreceptor tyrosine-based inhibitory motifs (ITIMs) of PECAM-1. To date, the mechanism of trans-homophilic interaction of PECAM-1 remains unclear. Here, we present the crystal structure of PECAM-1 IgL1-2 trans-homo dimer. Both IgL 1 and 2 adopt the classical Ig domain conformation comprised of two layers of β-sheets possessing antiparallel β-strands with each being anchored by a pair of cysteines forming a disulfide bond. The dimer interface includes hydrophobic and hydrophilic interactions. The Small-Angle X-ray Scattering (SAXS) envelope of PECAM-1 IgL1-6 supported such a dimer formation in solution. Cell adhesion assays on wildtype and mutant PECAM-1 further characterized the structural determinants in cell junction and communication.
- School of Biomedical Sciences, University of Hong Kong, Laboratory Block, 21 Sassoon Road, Pokfulam, Hong Kong, China.
Organizational Affiliation: